Research Summary
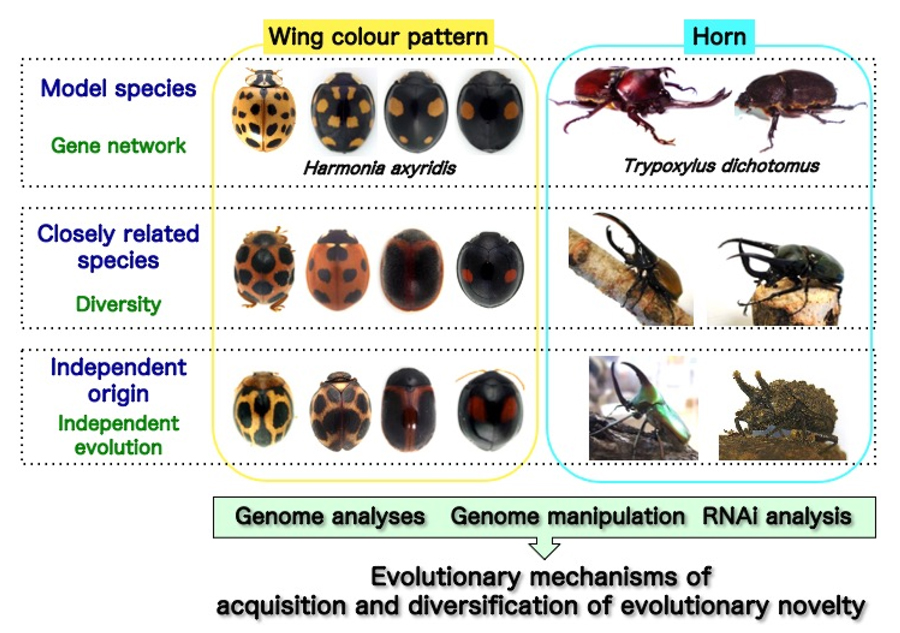
We focus on the evolutionary novelties acquired by insects in order to elucidate the molecular and evolutionary mechanisms that lead to the large variety of traits that they display. Among this wealth of exciting traits, our lab is currently focused on promoting research into (1) the origin and diversification of insect wings, (2) wing color patterns and mimicry of ladybird beetles, and (3) the acquisition and diversification of beetle horns.
Research Projects
-
• Left-right asymmetry of paired organs in insects
-
• Wing color pattern formation in the ladybird beetle Harmonia axyridis
-
• Mimetic wing color pattern formation in the flea beetle Argopistes coccinelliformis
-
• Horn development in the Japanese rhinoceros beetle Trypoxylus dichotomus
-
• Establishment of systems for gene function analysis in non-model insects
I. Origin and diversification of insect wings
The flight organs of insects have uniquely evolved when compared to that of other various flying animals on earth. Despite more than two centuries of debate, the evolutionary origin of insect wings is still an enigma; one which we are trying to decipher by the use of evo-devo methods. In Drosophila melanogaster, the wing master gene vestigial (vg) and its interaction partner scalloped (sd) play pivotal roles in the formation of wing field identity. For this reason, these genes are ideal research candidates in the investigation of wing origin and evolution.
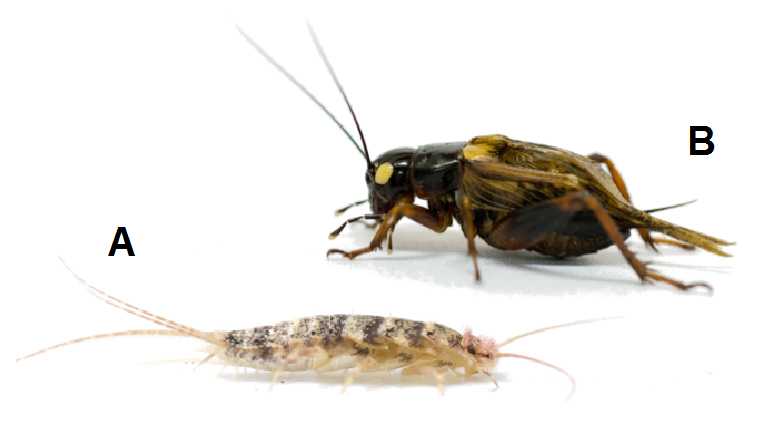

Figure 1. The firebrat,
Thermobia domestica (A). The two-spotted cricket,
Gryllus bimaculatus (B).
One way to identify the structure from which insect wings first evolved is to explore the function of “wing” genes in ancestral wingless (apterygote) species. To achieve this end, we chose the firebrat, Thermobia domestica, as a model (Figure 1A). T. domestica belongs to Thysanura, which is phylogenetically the closest extant relative of winged (pterygote) insects, thus making it ideal for elucidating wing origin. We cloned vg and sd orthologs from T. domestica (Td-vg and Td-sd), and developed RNA interference (RNAi) based methods for T. domestica to examine the functions of these genes. We are currently testing the functional effects of altered transcription for each of these wing genes in ancestrally wingless firebrats. Furthermore, we are performing comparative analyses of the function of these same genes in “primitively winged” (hemimetabolous) insects (Figure 1B) to obtain additional clues relevant to us understanding the origin and evolution of insect wings.
Interestingly, our previous work showed that vg expressing epidermal tissue forms lateral outgrowths in non-winged segments in the mealworm beetle (Ohde et al., 2013). Based on this, we hypothesize that ancestral lateral body wall outgrowths evolved into functional wings. However, genetic tools available for the analysis of basally branching wingless species are limited. To overcome these limitations, we established CRISPR/Cas9-based germline genome editing in T. domestica. Heritable mutations were successfully introduced in white locus, an evolutionarily conserved gene, encoding the ATP-binding cassette (ABC) membrane transporter, of T. domestica by using CRISPR/Cas9 system. This in turn results in white-eyed firebrats. In addition to the RNAi-mediated gene knockdown (Ohde et al., 2009), germline genome editing using CRISPR/Cas9 in T. domestica provides a platform technology for creating new research opportunities concerning the evolution of insects, such as insect wing origin. We are now conducting gene knock-out/in within various “wing” genes to identify genetic details and cell lineage analyses in T. domestica (Figure 1).
II. Wing color patterns and mimicry of ladybird beetles
A tremendous range of diversity in wing color patterns has evolved among insects, which in turn plays various ecologically important roles such as intraspecific sexual signaling, mimesis, mimicry, and is also used as a warning signal to predators. However, the molecular mechanisms responsible for generating such color patterns in most ladybird species remain elusive. To investigate the developmental mechanisms of color pattern formation, we have been focusing on the multicolored Asian ladybird beetle, Harmonia axyridis, which has conspicuous and variable wing color patterns consisting of black and red pigments (Figure 2A). The ladybird’s vivid wing color pattern acts as a warning signal to predators that they taste bad. At the same time, various other insect species utilize this ecological signal by mimicking the ladybirds’ wing color patterns. Mimicry provides us with an exciting opportunity to study how independent lineages of insects have evolved convergent color patterns. To explore color pattern formation mechanisms in mimicry, we are currently focusing on the leaf beetle, Argopistes coccinelliformis, which has color patterns similar to Harmonia, and is thought to be a Batesian mimicry of ladybird beetles (Figure 2B). To elucidate the molecular mechanisms underlying these wing color patterns, we established a technique for germline transformation using a piggyBac vector and RNAi in the ladybirds.
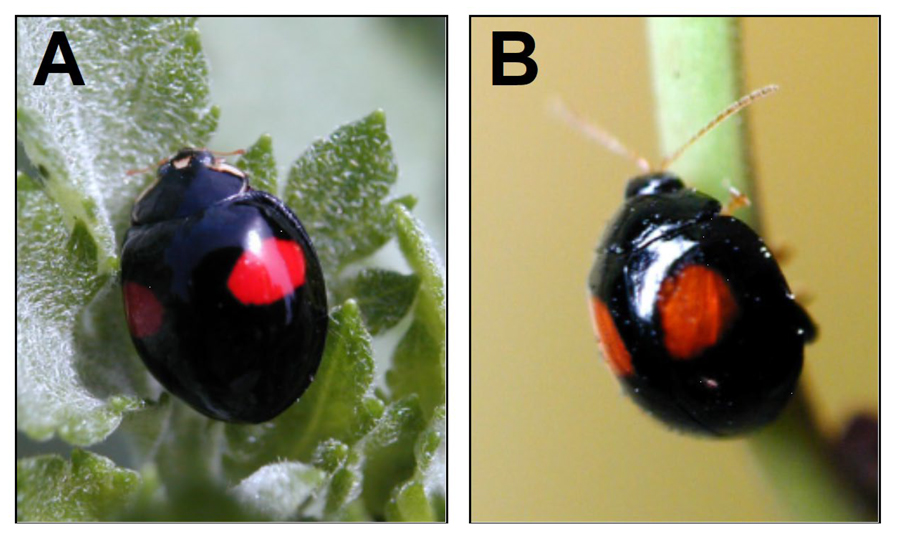

Figure 2. The ladybird beetle, Harmonia axyridis (A) and the leaf beetle, Argopistes coccinelliformis (B).
We recently identified a key gene, pannier, which regulates intraspecific color pattern polymorphism in H. axyridis using next generation sequencing technologies (RNA-seq and de novo genome assembly), and an RNAi-based screening method that we have established. pannier is expressed in specific regions in the wing, which synthesizes black pigment, and suppresses red pigmentation. The expression pattern of pannier is diversified according to the diverse color pattern types in H. axyridis. These findings suggest that regulatory shift, such as changes in enhancer activity, at the pannier locus may be crucial for the evolution of wing color patterns in H. axyridis. We are currently trying to elucidate the evolutionary origin of color patterns in ladybirds with a focus on regulatory shifts at the pannier loci.
We are also attempting to apply genome-editing technologies such as TALEN and CRISPR/Cas9 to tackle this issue. Thus far, we have achieved an efficient method of gene disruption using TALEN. Recently, we have begun successfully establishing the disrupting pannier gene function using CRISPR/Cas9 system. Ribonucleoprotein (RNP) complex of Cas9 protein and guide RNA targeting pannier coding exon was microinjected into the fertilized eggs. As a result, over 60% of G0 founders produced pannier knockout progenies (Figure 3). CRISPR/cas9 mediated gene disruption has advantages in terms of ease of designing gRNA and also possesses a high KO efficiency compared to TALEN mediated gene modification in Harmonia. To this end, we are continuing to develop CRISPR/Cas9 mediated gene knock-in method. To apply this approach, we are trying to attempt more complicated genome editing techniques such as genomic insertion, inversion and duplication to identify the crucial regulatory shift that may have driven the evolution of wing color patterns in ladybird beetles.
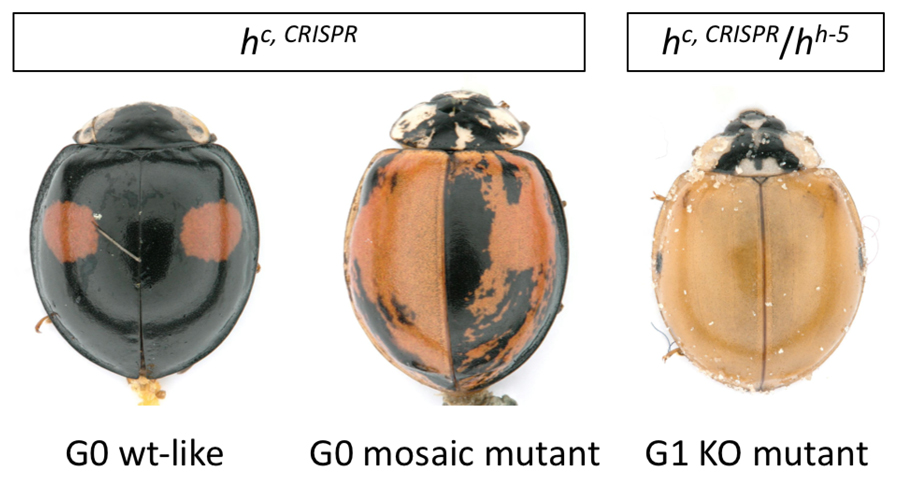

Figure 3. Representative photographs of pannier mutant phenotypes in G0 wt-like (left side), G0 mosaic (middle side) and G1 KO mutant (right side).
We are also establishing cryopreservation methods for germline cells in ladybird beetles to assist us with this. This is due to the high risk of losing valuable genetic bioresources in non-model insects. We recently established ovary transplantation and ovarian cryopreservation techniques in ladybird beetles. We hope that the genetic tools and techniques that we have established will further facilitate this research.
We plan to eventually analyze how the similar wing color patterns of model and mimic are generated based on the knowledge obtained from H. axyridis. For example, do they use conserved or divergent mechanisms?
III. Acquisition and diversification of beetle horns
Insects show a tremendous range of diversity in “horns”; rigid body outgrowths that function as weapons. Horns are a subject of great potential for evo-devo studies because they have arisen de novo multiple times as evolutionary “novelties”. However, the molecular mechanisms involved in sexually dimorphic horn formation are still poorly understood. To investigate the developmental mechanisms of horn formation, we are focusing on the Japanese rhinoceros beetle, Trypoxylus dichotomus (Coleoptera), which exhibits remarkable sexual dimorphisms in head and thoracic horns. The male-specific horns of T. dichotomus are among the best models for studying how an extreme, sex-specific morphological structure is formed (Figure 4 A).
We first investigated the process of horn primordium formation to understand the developmental mechanism of horn formation. The insect body is covered by a rigid exoskeleton mainly consisting of chitin, so changes in body size and shapes are produced by molting. Beetle horns first appear at the pupal period (Figure 4 A). During the prepupal period (about 130 hours) which is an intermediate stage between the last larval instar stage and pupation, the head horn primordia are packed folded inside the larval head capsule (Figure 4 B). This folded horn primordium develops into its final 3D shape (pupal horn) during pupation. To properly make the horn during pupation, the folding pattern (Figure 4 B) and surface micro furrows (Figure 4 C) are precisely formed on the horn primordium and packed within the confined space of the prepupal head. We recently attempt to use micro-CT to analyze 3D pattern formation of head horn primordia during the prepupal period. We found that horn 3D morphogenesis is accomplished in only 72 hours from the prepupal initiation (Figure 4 B, C).
We are currently focusing on this developmental stage because crucial regulatory factors for horn formation and differentiation are supposed to be activated at this stage in T. dichotomus. The present study provides a good starting point to unveil the gene regulatory network for sexually dimorphic horn formation and to pursue the evolutionary origin of such a regulatory system.
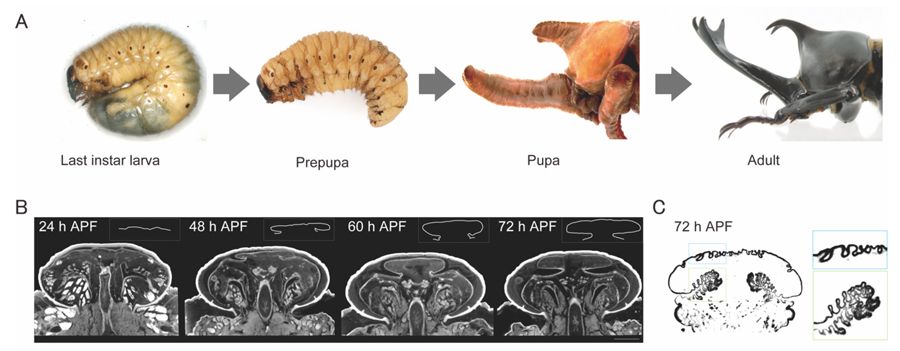
Figure 4 Morphological change of horn primordia in Trypoxylus dichotomus.
(A) Developmental process of Trypoxylus dichotomus male. (B, C) Micro-CT images of the male head horn primordium during pupal development. (Adapted from Morita and Niimi, Sanshi Konchu Biotec, 86: 145–151, 2020.)
Selected Publications
Morita, S., Shibata, T.F., Nishiyama, T., Kobayashi, Y., Yamaguchi, K., Toga, K., Ohde, T., Gotoh, H., Kojima, T., Weber, J.N., Salvemini, M., Bino, T., Mase, M., Nakata, M., Mori, T., Mori, S., Cornette, R., Sakura, K., Lavine, L.C., Emlen, D.J., *Niimi, T., and *Shigenobu, S. (2023). The draft genome sequence of the Japanese rhinoceros beetle Trypoxylus dichotomus septentrionalis towards an understanding of horn formation. Sci. Rep. 13, 8735.
Chikami, Y., Okuno, M., Toyoda, A., Itoh, T., and Niimi, T. (2022). Evolutionary history of sexual differentiation mechanism in insects. Mol. Biol. Evol. 39, msac145.
Sakai, H., Oshima, H., Yuri, K., Gotoh, H., Daimon, T., Yaginuma, T., Sahara, K., and Niimi, T. (2019). Dimorphic sperm formation by Sex-lethal. Proc. Natl. Acad. Sci. USA 116, 10412-10417.
Morita, S., And, T., Maeno, A., Mizutani, T., Mase, M., Shigenobu, S., and Niimi, T. (2019). Precise staging of beetle horn formation in Trypoxylus dichotomus reveals the pleiotropic roles of doublesex depending on the spatiotemporal developmental contexts. PLoS Genet. 15, e1008063.
Ohde, T., Morita, S., Shigenobu, S., Morita, J., Mizutani, T., Gotoh, H., Zinna, R.A., Nakata, M., Ito, Y., Wada, K., Kitano, Y., Yuzaki, K., Toga, K., Mase, M., Kadota, K., Rushe, J., Lavine, L.C., Emlen, D.J., and Niimi, T. (2018). Rhinoceros beetle horn development reveals deep parallels with dung beetles. PLoS Genet. 14, e1007651.
Ando, T., Matsuda, T., Goto, K., Hara, K., Ito, A., Hirata, J., Yatomi, J., Kajitani, R., Okuno, M., Yamaguchi, K., Kobayashi, M., Takano, T., Minakuchi, Y., Seki, M., Suzuki, Y., Yano, K., Itoh, T., Shigenobu, S., Toyoda, A., and Niimi, T. (2018). Repeated inversions within a pannier intron drive diversification of intraspecific colour patterns of ladybird beetles. Nat. Commun. 9, 3843.