Basic Biology Program, SOKENDAI (the Graduate University for Advanced Studies)
NIBB Academic Research Experience Program
(formerly known as the NIBB Internship Program)
Message from Interns 2024

Hanoi University of Science, Vietnam
Ha Linh Tran
My name is Linh, an undergraduate student from Hanoi University of Science, and I am currently working at the Vietnam Academy of Science and Technology. I had a wonderful opportunity to join the NIBB Internship Program in 2024 and spent my internship in the lab of Professor Morita. The one-month internship provided me with invaluable experiences—not only did I acquire new knowledge and lab skills, but I also learned a great deal about work organization, data analysis, and scientific methodology. Moreover, it was truly an eye-opening experience. The working style, mindset, and passionate attitude of everyone in the lab showed me what real scientific research is like. It made me realize that I still have a great passion for research and aspire to pursue it in the future, especially in an international research environment. Moreover, what impressed me the most was the warm welcome from all the lab members. They were always willing to help me, not only with my work but also in my daily life throughout my time in Japan.
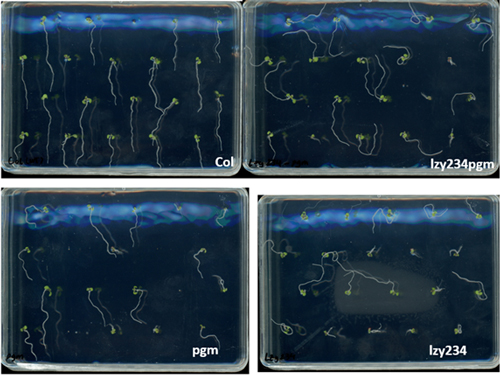
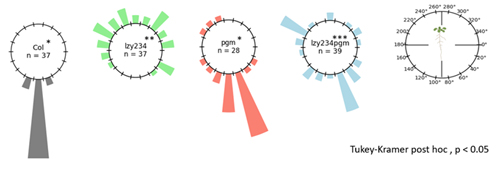
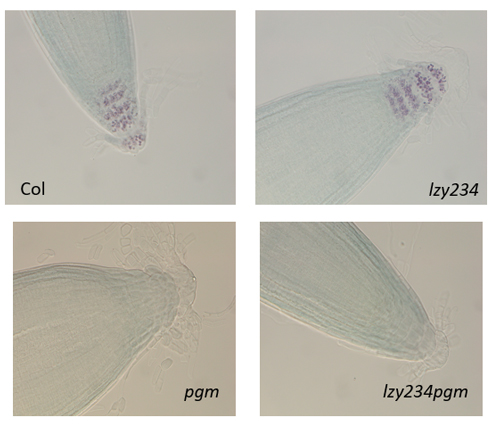
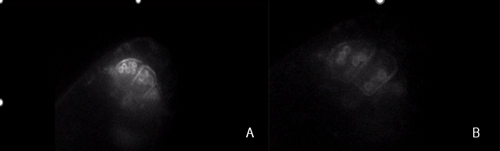
This was truly a wonderful research opportunity where I had the chance to explore and gain valuable insights into gravitropism, an intriguing response mechanism in plants that I had previously only learned about in school. For the first time, I delved into the molecular mechanisms, potential protein families, and their possible interactions that drive plant responses to gravity. Under the guidance of Prof. Morita, Dr. Nishimura, Dr. Kawamoto, Dr. Shikata, and Dr. Mori, I acquired many new skills and knowledge. From this experience, I developed a comprehensive understanding of the basic step-by-step procedures and learned how to conduct research on gravitropism using Arabidopsis as the model organism. I participated in four main research objectives. First, I learned how to evaluate the differences in phenotypes of some mutant lines and compare them to the wild-type (Col) to assess the impact of these mutations on plant morphology and predict the roles of the corresponding genes. Through this, I became familiar with key steps such as medium preparation, sowing seeds, growing plants in soil, and phenotype evaluation methods such as starch staining and measuring root tip angles. The three mutant lines I analyzed were pgm (a mutation in the gene encoding PGM-Phosphoglucomutase), lzy234 (a LAZY1-LIKE (LZY) triple mutant), and lzy234pgm. The evaluation results showed that the lzy234 mutation severely impairs the gravitropic response in roots (causing most roots to grow upward instead of following the gravitational direction), but it does not affect starch synthesis or sedimentation (as indicated by the iodine staining results at the root tips of these mutants). This suggests that the LZY protein could play a vital role in auxin polarization or other downstream signaling processes. Although PGM is involved in the interconversion of glucose-1-phosphate (G1P) and glucose-6-phosphate (G6P), a crucial step in starch synthesis, the gravitropic deficiency of pgm and lzy234pgm was less severe than that caused by lzy234. This finding suggests that mechanisms beyond the starch-statolith hypothesis might contribute to gravity sensing and root orientation, and that pgm could interfere with the impact of lzy234. Second, I learned how to select the desired transgenic lines (homozygous in transgenic genes) by analyzing the segregation ratio, which truly impressed me because it was the first time I had the chance to apply Mendel’s Law in such a clear and practical way. Additionally, I learned how to select homozygous mutants in RCC1-like domain (RLD) proteins—specifically the rld triple mutant—using genotyping. The main objective was to obtain rld123 expressing PIN2-GFP and ARF1-GFP to observe the localization of these two proteins in the triple mutants. Lastly, I performed the Yeast Two-Hybrid System for the first time to detect the interaction between the LZY1 protein and the RLD1 protein, particularly between the CCL domain (LZY) and the BRX domain (RLD). The process of conducting the experiment and evaluating potential sources of error allowed me to develop deeper problem-analysis skills and draw valuable lessons for myself. Moreover, I had the opportunity to learn how to use various types of microscopes, ranging from basic to highly advanced ones—some of which I had never seen before. All of them were fascinating and incredibly exciting. Using these microscopes, I was able to observe asymmetric root growth following the gravity direction, as well as the relocalization of LZY4-mScarlet when altering the gravity direction.
In conclusion, this has truly been the most incredible experience I have ever had. Beyond gaining new knowledge and skills, I also learned so much about how scientific research is conducted. The lab presentations provided me with deeper insights into gravitropism as well as the work of other lab members, giving me a broader perspective on the responsibilities of PhD students in the lab. Everyone in the lab was incredibly friendly and patient with me, always willing to teach me step by step. They even introduced me to Japanese specialties and shared fascinating aspects of Japanese culture with me. Additionally, I had so many exciting experiences and trips in Japan—ones I never thought I would have the chance to see and try. I am truly grateful to Professor Morita, all the lab members, and the NIBB Committee for giving me this amazing opportunity. This is an experience I will never forget.

Heidelberg University, Germany
Tiago Cordeiro da Trindade
My name is Tiago Cordeiro da Trindade, and I am currently a first-year master student majoring in Developmental and Stem Cell Biology at Heidelberg University. I was honored to spend two months as an intern at the National Institute for Basic Biology (NIBB) in Okazaki, Japan, working under the guidance of Professor Kamei and Nonaka and their dedicated teams. This internship was a transformative experience that deepened my understanding of developmental biology and molecular regulation.
During my time at NIBB, I engaged in several projects that allowed me to explore techniques in gene editing and embryonic development. One of the primary projects involved inducing somite-specific gene editing in medaka embryos using the Gaudí toolkit and the IR-LEGO system. The objective was to achieve targeted heat shocks to induce Cre-LoxP recombination, enabling visualization of gene expression changes in specific somites.
I began by creating an F1 line of medaka embryos from a Cre driver line under a heat-shock promoter and a reporter line containing fluorescent proteins flanked by LoxP sites. This setup allowed for the observation of Cre-mediated recombination through the switching of fluorescent markers upon activation. Using the IR-LEGO system, I performed localized heat shocks on specific somites, which successfully resulted in targeted fluorescent signals (Fig. 1). Observing the precise expression of fluorescent markers in the somite cells was incredibly rewarding and affirmed the effectiveness of the techniques employed.
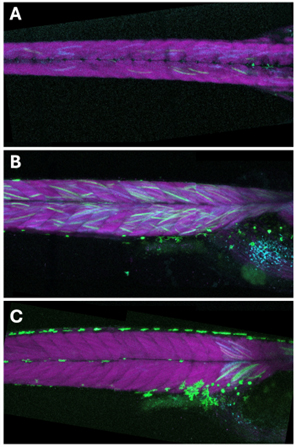
Another significant project was replicating the flow chamber experiment by Nonaka et al. to investigate left-right asymmetry in mouse embryos. The goal was to apply artificial rightward flow to pre-somite stage mouse embryos to see if it would reverse the usual left-right patterning. We prepared wild-type ICR mouse embryos and placed them in a flow chamber with the correct node orientation. Despite the meticulous setup and execution, the embryos developed with normal situs solitus. This outcome suggested that left-right determination might have occurred before the experiment, providing insights for future research.
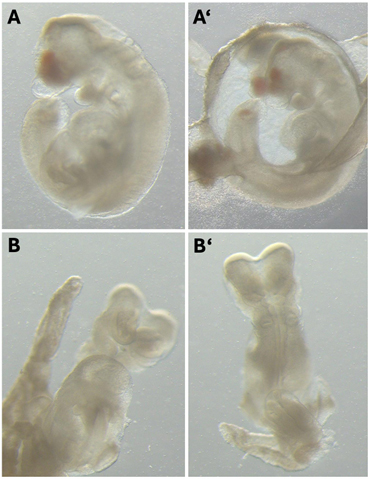
Additionally, I worked on transfecting NIH3T3 cells with the B-gTemp thermosensor to evaluate the effects of different promoters on protein expression and cell behavior.
Throughout these projects, I gained hands-on experience with advanced laboratory techniques such as microinjections, flow chamber setups, transfection methods, and fluorescence microscopy. I also developed critical thinking and problem-solving skills, especially when troubleshooting experimental challenges. Collaborating with the talented researchers and fellow interns at NIBB fostered a supportive learning environment that enhanced my scientific knowledge and interpersonal skills.
Beyond the laboratory, immersing myself in Japanese culture was an enriching aspect of my internship. Exploring Okazaki, participating in local traditions, and building friendships with lab members made my stay memorable. The hospitality and kindness I encountered left a lasting impression on me.
I am profoundly grateful to Professor Kamei, Nonaka, Yoke, Hayashi, Okada and all the members of the laboratory for their mentorship and encouragement. Their expertise and patience were invaluable to my growth as an aspiring scientist. I also want to extend my sincere appreciation to the NIBB Internship Program for providing this opportunity.
The knowledge and experiences I gained at NIBB have inspired me to pursue further studies and contribute to the scientific community. I look forward to building upon this foundation in my future academic and professional endeavors.

Guru Gobind Singh Indraprastha University, New Delhi, India
Tanya Varma
My name is Tanya, and I am a postgraduate from Guru Gobind Singh Indraprastha University, New Delhi, India. Through the NIBB internship program, I had the privilege of joining Nakayama Sensei’s lab as an intern from September to December 2024. This experience was transformative, offering invaluable insights into advanced molecular biology techniques and experimental approaches.
During my internship, I focused on purifying subunit 2 of RNA Polymerase II, gaining hands-on expertise in techniques such as Tandem Affinity Purification and Western Blotting. These tasks honed my skills in sample preparation, protocol execution, and data analysis. The opportunity to work closely on protein purification and expression studies not only enhanced my technical proficiency but also deepened my understanding of the intricate processes involved in gene cloning.
Beyond the lab, the internship allowed me to immerse myself in Japan’s rich cultural tapestry. I had the chance to explore traditional customs, savor diverse cuisines, and visit various regions, all while experiencing the warmth and hospitality that made me feel like part of the community. This cultural exposure beautifully complemented my academic pursuits, enriching my overall experience.
The supportive and collaborative environment in Nakayama Sensei’s lab played a crucial role in making this journey memorable and impactful. I am profoundly grateful to Nakayama Sensei and the entire lab team for their guidance and encouragement, as well as to the NIBB program for facilitating this incredible opportunity. This internship not only broadened my academic and cultural horizons but also reshaped my approach to research and strengthened my passion for scientific discovery.
Message from Interns 2023

Fulbright University Vietnam Vietnam
Nguyen Duy Hieu
Currently, I am a fourth-year undergraduate student majoring in Biological and Health Sciences, with a minor in Linguistic Anthropology at a small liberal arts college in Vietnam. In summer 2023, I had a great honor to join Prof. AOKI’s lab as an internship student for the NIBB internship program 2023. During my 10-week stay, I had one of the best times for learning, living and truly doing science, for which I was able to grow not just as an aspiring scientist, but also as a person. Everyone in AOKI lab never stopped amazed me with their warm and kind hospitality, their great passion and excellence in doing research. This experience had taught me some invaluable lessons, and fostered some meaningful connections that I would like to cherish!
What I learn and appreciate the most is the true scientific spirit that everyone in the lab cultivates. Under the guidance of Dr. SUGIYAMA, Dr. GOTO, and Prof. AOKI, I was trained to ask critical questions for my research, with a healthy and respectful skepticism. I have learnt to pay great attention and care to my works. They taught me from the smallest details, such as efficient pipetting skills or labeling my samples, to the most general concepts about science, academia, and life. I learnt the most basic to the cutting-edged techniques in biology, from simple gel electrophoresis or Western blot to complicated cell imaging processes. This internship has shown me great reflections about my research career. I think I have made the right choice, and now I am more than ready to pursue a scientific career ahead.
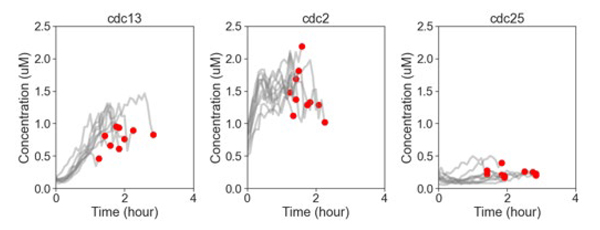
One of the main projects in which I was involved was to study the effects of nutrient conditions on division and morphology of S.pombe. To do that, I conducted time-lapse experiments of fission yeasts, whose CDK activity (i.e., phosphatase and dephosphatase activity) was measured by a newly developed FRET, in 02 nutrient conditions (i.e., rich vs poor in Carbon and Nitrogen source). I experimented on 2 yeast strains that were previously designed by Dr. SUGIYAMA. Fission yeasts in each of these strains were tagged with other phase-marker proteins (i.e., PCNA, Pom1, and Cdc13). This system allowed me to quantify the temporal dynamics between CDK activity and other related proteins involved in the governance of cell division cycle.
The results we found provided preliminary evidence that nutrient conditions did modulate changes in the cell cycle progression, reflected via (1) cell cycle duration, (2) CDK peak activity, (3) S-phase duration (Figure 1: Top, Middle, Bottom, respectively). In that, although the overall cell cycle duration of fission yeast in Poor condition was longer than that of Rich condition, the duration of S-phase was significantly extended in the latter condition. It was possible that S-phase in Poor condition was shorter due to the lack of nutrients, so that G1-phase, possibly G2-phase, was elongated as a compensating mechanism in Poor condition. In addition, we also found some complex changes in the morphology of Pom1-deleted cells, in response to the nutrient conditions. The clear relationships and mechanisms await further in-depth studies and analyses.
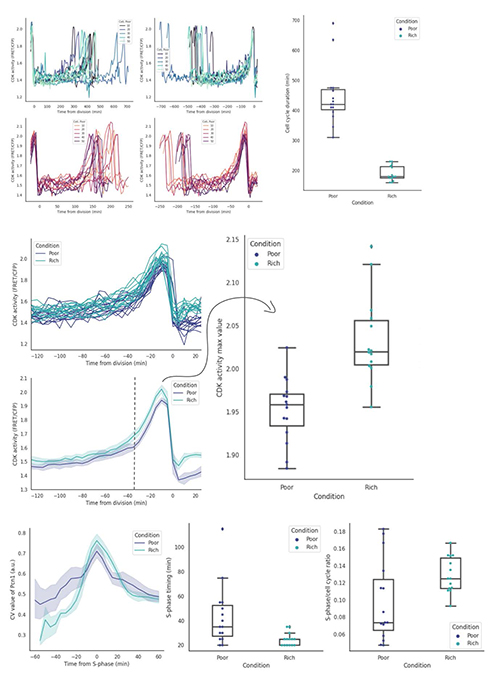
To see the cells progress through their cycle, with different protein expressions tagged with many fluorescent proteins, and to be able to analyze the results are truly fascinating to me. Additionally, I also learnt to clone DNA and design plasmid to create a new S.pombe strain whose heterochromatin assembly can be optogenetically manipulated. I was also able to participate in one small optogenetic experiment where the ZO-1 protein of HeLa cells was turned on and off by different lights. I literally wow-ed when the protein translocated to the cell’s membrane immediately after the light was shined onto the HeLa cells. The response was just in a blink of an eye. It was just WOW!
Once again, I am truly grateful for the opportunity. I would like to express my sincere thanks to everyone in AOKI lab. My time there was great, and I am missing you all already!

Kyushu University, Japan
Trang Thu Dao
In the scope of epigenetic changes, environmental stress leads to modifications to the DNA molecule or associated proteins that can be passed on to subsequent generations without altering the underlying DNA sequence. One of the major epigenetic information carriers is heterochromatin. In the case of heterochromatin components, the methylation status of H3K9 is crucial in epigenetics, regulating gene expression to cope with environmental stress. If a gene is turned off through histone modifications in a parent cell, its offspring cells might inherit the same gene-silencing pattern. Therefore, the big question is “How do histone modifications cause epigenetic inheritance for the next generation?”
Analysis of a mechanism of transgenerational epigenetic inheritance is the key to answering this question. During my time at the Division of Chromatin Regulation, I joined a project to integrate a tethering system into the genome of fission yeast Schizosaccharomyces pombe (S. pombe) by CRISPR/Cas9 system. The tethering system regulating genomic activity will be expressed during meiosis to study the mechanism of transgenerational epigenetic inheritance.
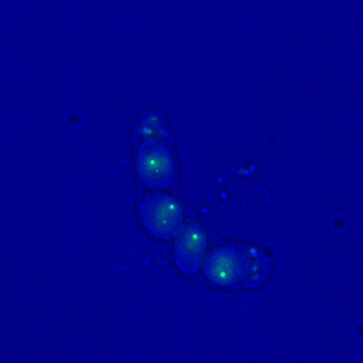
I constructed the pCas9-gRNA plasmid to create a DNA double-strand break for insertion by homologous recombination. After preparing donor DNA fragments for integration, I performed genome editing on S. pombe to insert the gene encoding the tethering system. I also analyzed the localization of telomeres in fission yeast spores under the microscope. As Taz1 is a subunit of the telomeric complex, it is used to fuse the GFP protein for localization. The telomeres were observed inside the fission yeast spores.
Along with learning new experimental techniques, I had opportunities to attend Joint Progress seminars, in which I could learn more new concepts and insights about histone modifications. The final presentation equipped me with the experience of professional scientific communication. I am very grateful for all the invaluable experiences in NIBB.
After all, I would like to express my appreciation to Prof. Nakayama, Dr. Hayashi, and all the members of the Division of Chromatin Regulation for their warm welcome and wholehearted support to me during my internship. I would like to extend my special thanks to Dr. Kiyoshi Tatematsu for his support to prepare administration documents. Last but not least, I also want to express my gratitude to the NIBB Internship Program for giving me this chance to experience research and life in Okazaki City.

Hanoi University of Science, Vietnam National University, Vietnam
Nguyen Ha Phuong Thao
(a)
(b)


Gyeongsang National University, Korea
Yeongcheon, Yang
It is true that my research in NIBB internship program mainly about the gravitropism, but I was possible to learn fundamental biological experiments from the selection process of mutants, which are the basis for research, to the interpretation of data. During the 3 weeks internship, I focused on three major things. First, selection of mutant plants. Second, relationship of LZY and Amyloplast. LZY is a key in gravity response of plant, and LZY stick to amyloplast’s outside surface and during the gravity stimulus move to plasma membrane from there. Lastly, the function of Amyloplast and LZY. Each LZY2, LZY3, LZY4 and PGM function was inferred from the phenotypes of single and double, triple, and quadruple variants.
To select specific mutant plants for research, I used antibiotic resistance selection (Kanamycin, Hygromycin B) and fluorescence methods. In this process, I was very surprised to find that Mendel's genetic laws, which were only seen in books, were very useful.
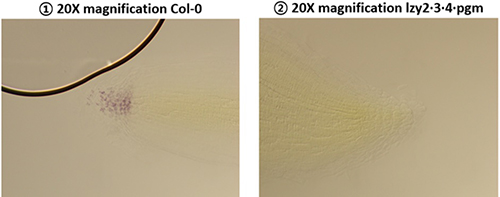
To identify phenomena occurring in columella cells during gravitational reactions, LZY4 protein containing mScarlet fluorescent protein was constructed (LZY4p:LZY4-mScarlet/lzy4). Plus, in order to compare the polar movements of LZY between normal and abnormal amyloplast, (LZY4p:LZY4-mScarlet/lzy4·pgm) was constructed. pgm, a key enzyme for starch synthesis, is essential for Amyloplast sedimentation and redistribution. Consequently, the pgm variant showed a large difference compared to plants with successful amyloplast precipitation and distribution of LZY4.
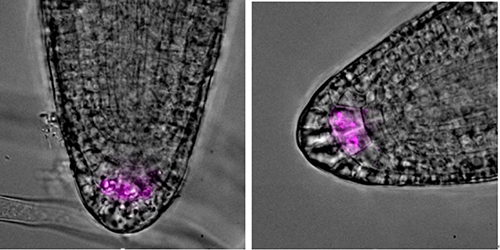
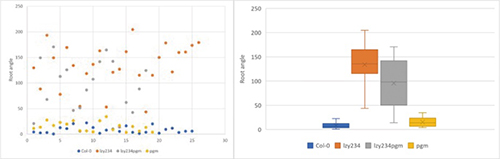
To examine LZY and Amyloplast properties, the root angles of each variant were calculated and starch in Amyloplast stained. In conclusion, Col-0 and pgm variant that have insufficient amyloplast sedimentation both show downward extension of main root and lateral root. The starch of Col-0 in columella cell was stained with 4% Idoine solution. contrastively, pgm knock out mutants doesn’t show stained starch.

Through the function of LZY and the indirect loss of amyloplast sedimentation of pgm mutants, I could guess the function of LZY affects to downward root growth. Furthermore, I could understand LZY plays a pivotal role in gravitropism more than I thought.

Overall, NIBB internship program not just helped me to learn a new concept, also to understand how valuable basic knowledge and methods is. I appreciate to all members of Prof. Dr. Morita laboratory, other technicians, and responsible people of this program. I am very grateful to them for providing to me precious opportunity to be in this program in Japan. Of course, I hope to back to Japan in the future, as well.

Hanoi University of Science and Technology, Vietnam
Nguyen Yen Chi
My project centered on Regiella insecticola, a facultative endosymbiont of aphids. Among the three common secondary symbionts in pea aphids – Hamiltonella defensa, Serratia symbiotica, and Regiella insecticola – the last one has been less extensively studied. Hence, my objective was to broaden the understanding of R. insecticola, particularly concerning its localization, transmission, and interactions with both the primary symbiont, Buchnera aphidicola, and the host insect. This was the first time I have ever worked with insects, offering me numerous novel experiences.
Initially, I detected the presence of this bacterium in several pea aphid strains in our lab through diagnostic PCR. Using fluorescent staining and FISH techniques, I observed numerous cells and organs in both viviparous and oviparous female individuals. The resulting images helped me localize the secondary symbiont and uncovered several findings regarding the transmission of R. insecticola throughout sexual and asexual generations. There were both novel observations and results that aligned with previous descriptions about other secondary symbionts. In the strain of pea aphid I studied, Regiella resides not only within the sheath cells but also outside of these cells and the bacteriocytes. Additionally, they were observed surrounding the fat cells and present inside some fat cells.
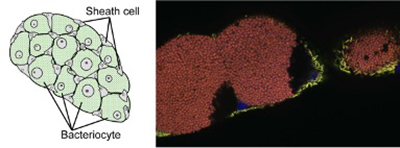
In my exploration of Regiella's characteristics and interactions, I familiarized myself with numerous command-line tools such as seqkit and local BLAST, conducted KEGG analysis, and utilized IGV for genome visualization. My skill set was expanded beyond the tools learnt in my bachelor's program. Additionally, I had the chance to take a Bioinformatics course in November, which provided an enlightening overview of the field through hands-on experiences. For me, this subject was new and challenging, and I received a lot of help from my lab members. As a result of new bioinformatics knowledge, I could discover new characteristics regarding the affection of Regiella to pea aphid and Buchnera. For example, it appears that Regiella does not fully compensate for Buchnera in many amino acid biosyntheses. In the specific strain that I studied, Regiella seemed to have positive affection on the growth speed of host aphids.
Throughout my internship at Shigenobu Lab, I was fortunate to receive invaluable guidance and support from all lab members. I have admired and learned a great deal from their work ethics, their knowledge, and their broad perspectives. I owe a debt of gratitude to Dr. Tomonari Nozaki, our lab's assistant professor, for his guidance in my lab work as well as his assistance in my social life. Beyond a mentor, he and his family are precious friends of mine. Overall, I genuinely feel blessed to have crossed paths with such exceptional people.
For all these priceless experiences, I am incredibly grateful to Professor Shuji Shigenobu and NIBB Internship Committee for accepting me to the Internship Program 2023. This experience allowed me to work and learn under Professor Shigenobu's guidance in his laboratory, granting me a broad range of knowledge and practical skills in an entirely new field of study. Furthermore, I've had the honor of meeting intelligent, kind-hearted people and building lasting friendships. This time become an invaluable journey filled with learning, personal growth, and unforgettable memories that will always be dear to me.

University of Heidelberg, Germany
Balkrishna Baral
My research primarily focused on utilizing Medaka and HeLa cells as models for studying different biological processes. For example, I utilized scanning electron microscopy (SEM) to visualize the micropyle in Medaka embryos. Additionally, I conducted experiments using the Gaudi (Hsp-Cre) * TG861(bactin-loxp-DsRed2-loxp-GFP) Medaka line. In response to IR LEGO, the region originally expressing DsRed2 turned green upon Cre induction, illustrating the dynamic changes in gene expression.
Beyond the laboratory, my internship experience extended to exploring the rich cultural tapestry of Japan. I had the opportunity to immerse myself in different cultural traditions, savor diverse cuisines, and explore other regions. The warmth and hospitality I experienced made me feel like part of the family. The supportive lab environment and cultural exposure facilitated a well-rounded learning experience.
In conclusion, my internship at Kamei Sensei's lab within the Optics and Imaging Facility was an enriching journey that reshaped my approach to science and broadened my horizons both academically and personally. I am immensely grateful for the opportunity and the invaluable lessons learned, which will undoubtedly guide my future research endeavors. I am thankful to Kamei Sensei and the entire lab team for their guidance and support throughout my internship and to the NIBB program for facilitating the internship.

University of Chinese Academy of Sciences, China
Kang Lyu
During my internship, I was mainly engaged in exploring the regulatory mechanism of Chlamydomonas reinhardtii photoprotection. An earlier work in the Minagawa lab showed that Algal photoprotection is regulated by the E3 ligase CUL4–DDB1DET1. The putative E3 ubiquitin ligase CUL4–DDB1DET1 in unicellular photosynthetic organisms may mediate blue-light signals to influence photoprotective gene LHCSR1 and LHCSR3 expression, thus exercising the role of photoprotection regulation. Besides, another result revealed CONSTANS flowering complex in the green alga Chlamydomonas has an important function in controlling the response of photoprotection. On the basis of the above studies, we look for other possible mechanisms that regulate the response of photoprotection.
Last but not least, I would like to express my sincere thanks to Professor Minagawa and Dr. Kamada and Dr.Makio and Dr. Kim and Ms. Yuka and other members of the laboratory. I enjoyed the time in NIBB very much. Although the time was not long, I felt the ample care and concern in such a harmonious family and felt the charm of scientific research. I would also like to thank Dr. Tatematsu and the NIBB Office for their thoughtful assistance in visa processing and during the internship. I hope I will have another opportunity to pursue scientific exploration here in the future.
Message from Interns 2022

University of Michigan, USA
Zeeshaan Sultan
Having worked with the Aoki Lab for five weeks, I was able to become proficient in wet lab techniques including plasmid construction, transfection with LentiX-293T, infection of HeLa cells, and live cell imaging. Thanks to the exemplary teaching by Dr. Tany and Dr. Goto, I was able to successfully create fluorescent biosensors for Ca2+, RhoA, KTR, and cAMP. In previous experiments, a biosensor system for these intracellular GPCR ligand binding activation signals was created; Ca2+ by R-GECO1 (Red Fluorescence), cAMP by EPAC-FRET (Cyan-Yellow Fluorescence), ERK by ERK-KTR (Near Infrared), and RhoA by DORA-RhoA (Cyan-Yellow Fluorescence); however, there was a problem with utilizing the same Cyan-Yellow fluorescence for cAMP and RhoA.
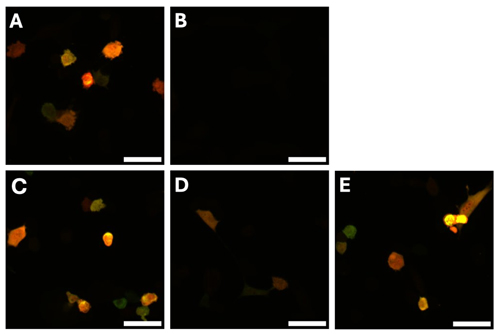
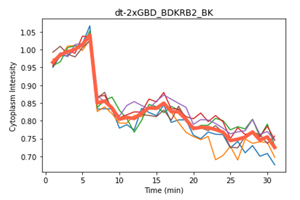
Together with the Aoki Lab members, we developed a new system for visualizing these intracellular signals that contains no overlapping colors; Ca2+ by B-GECO1 (Blue Fluorescence), cAMP by G-Flamp (Green Fluorescence), ERK by ERK-KTR (Near Infrared), and RhoA by dT-2xGBD (Red Fluorescence).
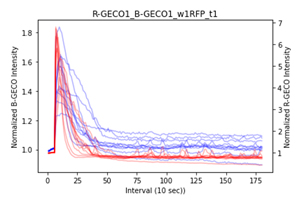
First, we wanted to confirm that B-GECO could be used as a Ca2+ sensor similar to R-GECO, and this was confirmed by the data analysis of co-cultured B-GECO and R-GECO cells. We found that B-GECO and R-GECO were similar as they both showed an initial peak for the influx of calcium (Fig 1.), and showed similar pulsing of cells. Next, we wanted to compare ERK-KTR-miRFP703 and ERK-KTR-iRFP, and the results were quite similar (Fig 2&3.); however, ERK-KTR-iRFP live cell imaging showed better differentiation of the nucleus and cytoplasm, so it was chosen as the better biosensor to be used in the bulk cell line.
Lastly, when developing a RhoA sensor, we decided to use dT-2xGBD as our biosensor of choice, as it was able to detect cytoplasm RhoA localization to the plasma membrane.
However, this was only possible in cells transfected and expressing the GPCR BDKRB2, a bradykinin receptor. The quantification of the live cell imaging of RhoA activation was interesting as it was able to show slivers of RhoA activation in the plasma membrane.
While the research was important, I also have fond memories of discussing a variety of old and new anime and manga with the other members of the lab. Every Thursday, I would go out to dinner with a few lab members and play Badminton with other research members afterwards. Although I was never very good at Badminton, I was able to meet new people and make new friends. I also was able to play soccer and explore Okazaki City and its numerous cultural and historical landmarks, and got better at Japanese due to conversing with the Aoki Lab members who were also able to help me study and become better at my Japanese grammar and increase my vocabulary.
I loved sampling the different Japanese, Chinese, and even Korean cuisine in the city of Okazaki; one of my recommendations would be the Japanese maze-soba or Nene Fried Chicken near the Higashi Okazaki Station. During the weekends, I enjoyed taking the train or Skinkansen and exploring the vast cities of Nagoya and Tokyo, which I highly recommend. Having a farewell celebration at the end of my time in the lab, I was able to present all my research throughout the internship, and am forever grateful for the opportunity to research with the Aoki Lab and would like to thank the NIBB Internship Program and Aoki Lab.

Mulawarman University, Indonesia
Stephani Layuk
During my internship period, I have worked on experiment about LHCBM1-4L mutant in Chlamydomonas reinhardtii to understand the function of amino acid in certain region of type IV Light Harvesting Complex II (LHCII) protein namely LHCBM1 from another previous experiment of this laboratory. Comparing to th other type of LHCII, LHCBM1 has a unique amino acid in N-terminal sequence that can not be found in other LHCBM. Several procedures has been carried out including culturing C. reinhradtii, performing PCR, made transformant, and running SDS-PAGE, which are some of the equipments were I newly used during my internship in this lab. Other than that, I really enjoyed my short-term stay in Okazaki. So many new fascinating experiences and places that I have visited in Japan. The accomodation is also convenient and provides many things that I needed.
Lastly, I would like to convey my gratitude to NIBB for giving me the opportunity as an internship student in one of their divisions. I also thankful for the heart-warming welcome from every lab members during my time in the lab, especially Prof. Minagawa as the head of the lab who had invited me in his lab and gave me precious advice on how to understand the advance research and study in the lab as well, for other lab members, Kamada-san, Kim-san, Kosugi-san, and everyone in the lab for welcoming me and help me out on understanding the research and also what I need during my stay in Okazaki, I could not thank enough for their kindness, actually. Hopefully, we could meet again in other occasion or somehow, I would have another chance to return back to NIBB.

Kyushu University, Japan
Muhammad Azka Aufa Ramadhani
During my visit, I was participating in an ongoing research project on endosymbiont bacteria of Pseudoregma panicola and Pseudoregma bambucicola aphid. These two species are close relative of a social aphid Ceraovacuna Japonica where a novel multi-partner symbiosis involving Buchnera sp. and Arsenophonus sp. was recently found. The target of the project was to sequence and identify the endosymbiont bacteria of Pseudoregma panicola and Pseudoregma bambucicola by using nanopore sequencing and bioinformatic analysis. In the process, I was helped by Shunta Yorimoto-san, a member of this lab.
The first thing I did was library preparation of isolated DNA sample for nanopore sequencing and loading of prepared library to nanopore sequencer. This is an important experience for me since it was my first time using a next generation DNA sequencing machine.
The next step is bioinformatic analysis of sequencing data. The analysis was performed through UNIX commands. Genome assembly, local BLAST, and online BLAST analysis was performed to identify the endosymbiont present in host aphid samples. This part of the project was particularly interesting for me since it was my first time using UNIX. Even though it was difficult to familiarize myself with the UNIX command, the detailed instructions and guidance from Prof. Shigenobu and Mr. Yorimoto really helped me in finishing the projects.
At the end of my visit, I also participated in the lab meeting and presented my analysis results. I am very grateful to experience all the laboratory experience from experiments to scientific presentation. I was surprised at how much I learned during my 2 weeks internship program.
Once again, I am really grateful for the opportunity to visit Prof. Shigenobu’s laboratory. Everyone in the laboratory was really kind and supportive on helping me who has almost zero bioinformatic experiences. Not only guiding me in performing experiments, the lab members were also kind enough to share their personal experiences in pursuing their education. They even gave me some advice on pursuing my scientific education.

Vietnam Research Station, Nagasaki University ‘s Project, Vietnam
Pham Hong Quynh Anh
During one month in Okazaki, I did learn how to effectively access bioinformatic tools and manage my database via VPN and BIAS system; and analyze gene ortholog-based sequence features via Microbial Genome Database (MBGD) which has been developing by Professor Uchiyama’s laboratory, by using my AMR samples collected from wastewater in Vietnam.
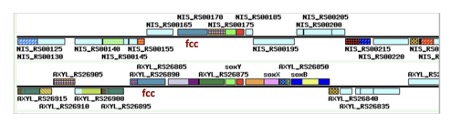
Orthologous clustering of all Aeromonas strains identified 7971 orthologous groups (OGs) defining the pan-genome. Histogram shows 2438 OGs which are conserved in all 21 strains, and represent the universal core among them. The remaining 5533 OGs correspond to the accessory genome, in which 173 OGs are unique in only NUITM-VA1. Compared to the reference, NUITM-VA1 strain lacks 7 proteins, which related to transport & binding of protein and transcription functions (Fig1).

Fig1. MBGD
Ortholog Cluster Overview.
Two NUTM-VA1 and NUTM-VA2 strains share 9 unique genes, which are absent in all Aeromonas strains in MBGD database. Besides, these strains were collected from urban drainage which located near some center hospitals in Hanoi, this clue suggests that these unique genes have some potential function to support bacteria survival in extreme environmental conditions (Table 1).
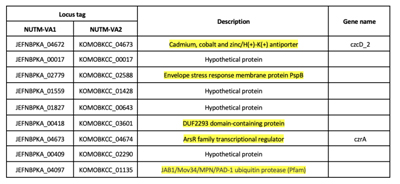
Table 1. Unique genes found in NUTM-VAs strains
Next, I conducted an experiment to identify horizontal gene transfer (HGT) acquired genes encoding proteins that function under particular conditions, such as genomic islands (GIs).
The results found 440 blocks related to mobility classes among 50 Aeromonas strains, partly shown in Fig2.
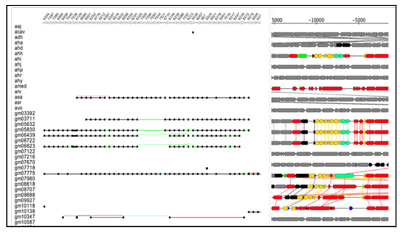

A black line represents a direct adjacency, a green line represents a non- adjacent neighborhood (i.e., there are insertions), and a red line represents an inversion.
The analysis conducted during this internship will be continued for the further analysis, to comprehensively answer my own question: “Which HGT and mechanism mainly affect the survival of bacteria in wastewater and resistant to antibiotics?”
In conclusion, NIBB Internship Program 2022 is an invaluable experience for me to consider entering the Ph.D. course in my next step career path. Despite the lack of time, I would like to express my sincere gratitude to Professor Ikuo Uchiyama, who accepted me to join his laboratory during my internship course. I wholeheartedly thank his patience, motivation and immense knowledge. Furthermore, I am thankful to Jiang-Kun, Kotani-san and her husband, for their kindly welcome to me to discover Okazaki sightseeing with historical views and yummy traditional foods. Special thanks to Prof. Kiyoshi Tatematsu for helping to prepare all documents. I also would like to send best regards to all staff in Uchiyama’s Laboratory and Mishima Lodge, who supported me during my one-month internship. Last but not least, many thanks are sent to my supervisors in Vietnam, my boyfriend in Tokyo and friends - alumni in NIBB, Okazaki, Aichi, Japan.

Vietnam National University, Vietnam
Pham Phuong Ngoc
My research topic radically focused on creating a stable HeLa cell line with 4 biosensors, detecting the fluctuations of cAMP existence, calcium concentration, RhoA translocation, and the dynamic of the ERK pathway through the activation of G protein-coupled receptor (GPCR).
Drowning in a sea of studies on GPCR, many cell cycle progressions, such as proliferation, differentiation, and cell survival, are shown to be related to a myriad of specific GPCR ligands. However, the mechanism of how signaling dynamics can quantitatively decide the cell's fate has yet to be unraveled. It is also noteworthy that cancer - a leading cause of death worldwide is basically the accumulation of cell cycle abnormalities. For this reason, we believe that visualizing the level of cell signaling after treating cells with different stimulants lays a concrete foundation for cancer research. Correspondingly, this experimental result may provide a good source of kinetic parameters for computer-assisted simulation, a potential approach for predicting the efficiency of medicine or drugs.
In 3 months, I dealt with DNA works to integrate the 3 types of fluorescence into 3 different and appropriate vectors. Then, the fluorescent expression vectors will be transferred into the HeLa cell line. After selecting cells with a vector of choice, fluorescent microscopy was subsequently utilized to examine the changes in fluorophore's color, depending on the cell signaling activities. Particularly, results obtained by time-lapse videos clearly showed me the colorful changes in a living cell. For me, that was incredibly awesome!
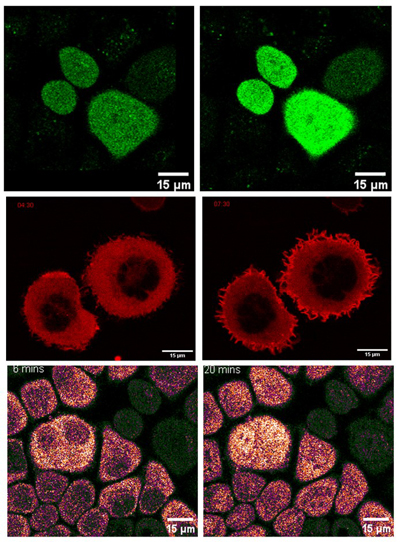
Note: (A) Activity of cAMP biosensor – GreenFalcan 0.3; (B) Activity of RhoA biosensor – dTomato; (C) Activity of ERK biosensor – KTR-iRFP.
During the time pursuing our main goals, I unintentionally made mistakes in experiments. Still, every time, Aoki professor and Aoki lab members were always there to give me a hand and, together with me fixing my problems. Through that, I have learned about so many things which are needed for working in a professional laboratory: carefully scheduling, fastly cracking mistakes, efficiently discussing with other members, and confidently giving and performing personal views of points.
Besides fulfilling my starvation for research with tons of invaluable lessons, I can also further my knowledge about Japanese traditional culture - my most interesting culture from the bottom of my heart since I already have a deep love for the 2D industry.
Touring the streets enveloped in sparkling lights, sampling many traditional delicacies such as Kirimochi or Kakumochi, Yakisoba, Karaage, Ramen, Red and White Miso, Oinari, Monaka, and many, many dishes that I had never tried before. In like manner, visiting a list of stunning temples, shrines, and the Kaiyukan aquarium in Osaka, entering Maid and Owl Coffee, trying the Game center in Osu Market, dipping in the yellow of autumn leaves in Kuragari and Korankei valley, enjoying the Halloween vibe under the Nagoya TV Tower, taking pictures with most creative cosplayers,... all distributed to my memorable moments here. And the most special thing that I cannot forget was my first time watching the Total Lunar Eclipse on November 7 and watching the Leonid Meteor shower at Otogawa river at 3A.M, then together with some of my precious Japanese friends making our wishes. That special day ended by flaking out at my friend's cozy house. We, 3 individuals, somehow slept in just a single bed and woke up at noon.
All in all, I would like to express my sincere thankfulness to my supervisor, Professor Aoki, and all laboratory members, who allowed me to practice and research at the Division of Quantitative Biology, National Institute for Basic Biology (NIBB). I really appreciate their support, guidance, understanding, belief, and, most importantly, encouragement. Consequently, I was trained on how to work/think critically and how to do research on my own.
Last but not least, with all regard and infinite gratitude, my thank would also send to NIBB for granting me a chance to work and study in such a wonderful workforce in my best living condition.
And my last words to all "greenhorns" like me from any parts of the world, I would highly recommend NIBB as your must-try stopover in your academic path and vigorous youth.

Kyungpook National University, Republic of Korea
Dang Huong Giang
The NIBB internship program is a wonderful journal. One week in Japan gave me the most memorable about the lab, the institute, as well as the culture and people that I cannot forget. I would like to express my gratitude to all members of the division of the embryology lab for your kind support and generosity not only in the lab but also in daily life. Moreover, the program gave me a chance to meet great friends. I want to send my thanks to all of them for their enthusiasm and friendly.
In the division of embryology lab, I had an opportunity to work with mass spectrometry (MS) of proteins of the embryo and the uterine lumen, and the microscope system for embryo time-lapse. The MS experiment followed embryo culture, embryo protein extraction with biotinylation, and Biotinylated peptide purification, and apply the sample to the MS device. The results of this experiment showed the list of proteins present in the embryo. The embryo time-lapse also started with the embryo culture, apply to the microscope, then keep watching them and analyzing the imaging. The video results can show the division and development process of the embryo directly. All of them is the new and unforgettable experiences for me. Furthermore, the one-week internship gave me many deep-knowledge of the field research in the lab and encourage me to be more hard-working and study.
The NIBB internship program gave me the opportunity to gain motivation and confidence for my future plans. Thank you so much for this chance and I hope to visit again in the near future.

Kyushu University, Japan
Yesaya Sumarli
Within the internship, I was given hands-on training on several experiments. The first and main experiment was CRISPR/Cas9 genome editing through microinjection on Medaka embryos. The target gene for knockout was SLC45a, which is associated with melanin formation. It was very challenging to handle and do microinjection on the embryos as it requires precise hand movements, and I failed multiple times at first. But I was given the time and materials to practice until I became more proficient with the technique. Several days after the microinjection, successfully genome edited embryos were identified by genotyping on resulting band separation of extracted embryo DNA in electrophoresis. In between those several days, I was also given training on medaka sperm collection and cryopreservation, as well as artificial insemination. I was told that Such techniques are really important in regard to preserve endangered or even extinct fish species. Furthermore, I got the chance to observe and even take a small part as a timekeeper on an experiment of an ongoing research of a neighboring laboratory.
Apart from practical experiment techniques, Prof. Naruse also gave me lectures about Medaka as a species, and their application as a model organism in experimental studies. Before coming to this internship, I never had any particular interest in Medaka. However, the lecture given by Prof. Naruse on Medaka combined with firsthand experience of its application, really opened my mind to the usefulness of Medaka for various research, and I became interested to work with it in the future.
To conclude, even though it was only for a short time, the internship program was a very fruitful and interesting experience to me. I would like to express my deepest gratitude to Prof Naruse and the staff in the Laboratory of Bioresource for training and taking care of me, as well as to NIBB for this internship opportunity.
Message from Interns 2021

Osaka University, Japan
Piyusha Mongia
The bigger question was to understand how epigenetic information is inherited and what are the factors involved. Epigenetics refers to heritable changes- not coded by DNA. Eukaryotic chromosomes can be divided into 2 broad categories: euchromatin and heterochromatin. Euchromatin refers to the regions on chromosomes which are transcriptionally active. Heterochromatin, on the hand, is generally transcriptionally silent, present at repetitive sequences or transposons. Methylation of lysine 9 of histone H3 (H3K9me) is a hallmark of heterochromatin. I created and assessed the involvement of four candidate gene deletions in heterochromatin inheritance in fission yeast spores. Fission yeast is a simple eukaryote having three chromosomes, making its study relatively easier. I first replaced the 4 genes with nourseothricin by PCR and then visualized swi6-egfp localization by fluorescence microscopy. Swi6 is a heterochromatin protein that is required for its assembly. One of the candidate genes seems to be important for the inheritance of heterochromatin but more studies need to be done to conclude so. In a parallel project, I tried to express three of Tetrahymena proteins using a bacterial expression system. These proteins, HPL4, HPL5, and JUB5 are involved in heterochromatin assembly. Producing recombinant proteins allows us to purify and biochemically analyze individual proteins activities and interactions in-vitro. In the end, I managed to express all, but purify one of the proteins due to time constraints. I learned many biochemical techniques, like Gel Filtration Chromatography and Ion Exchange Chromatography. I wish I could have had a longer time period to do more fun experiments.
Every day I was faced with challenges, and learnt something new. All the members were very patient with me and taught me something or the other. I’m extremely grateful to Dr. Hayashi and Dr. Kataoka, who guided me throughout the project. Prof. Nakayama was very supportive and encouraging. The research atmosphere in the lab is very exciting and it was truly an enriching experience for me.

Kyushu University, Japan
Geofanny Bernadette Yohanes
Although I did not participate in ongoing research, I was able to do three basic experiments. I did most experiments with the laboratory staff. The first was a knock-out of SLC45a, a gene controlling melanin synthesis, into medaka (Oryzias latipes). The second, also with medaka, was a knock-in experiment of mNeonGreen to the myosin heavy chain, creating a fluorescent fusion protein. Through both of these experiments, I learned and practiced techniques such as microinjection, DNA extraction, and PCR. At the end of each day, I was given a lecture by Prof. Naruse to prepare for the next step. This allowed me to gain a deeper understanding of knowledge I have learned in school, such as the mechanism of CRISPR/Cas9, differences between gene editing methods and when to apply them, and factors to consider in primer design. It was fun to watch the embryos grow and display the results of these experiments, although some must be used for genotyping to confirm the success. The third experiment was cryopreservation of medaka sperm and artificial insemination using the preserved sperm.
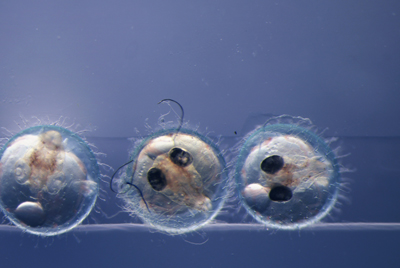
Result of knock-out experiment. Melanin formation in the eyes was affected.
In addition to learning knowledge and skills specific to each experiment, this internship gave me an insight on how to be a good researcher. I learned how important it is to keep an organized journal, to clean up and take care of your tools, and what discussion around research is like.
I would like to thank Prof. Naruse and the lab staff for hosting me during this internship. It has been an invaluable experience. The skills I learned will no doubt be useful for me as I plan to work in research related to fish in the future.

Kyushu University, Japan
Yiqi Yang
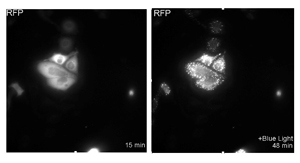
The purpose of my project is to verify the mitogen-activated protein kinase (MAPK) pathway by using two systems: optogenetics Cryptochrome-2 (CRY2) and Kinase Translocation Reporter (KTR). Upon photoexcitation (blue light), CRY2 proteins simultaneously oligomerize9 and bind to CIB1, which form clusters by promoting interconnection among CIB1-conjugated MPs (Figure 1).
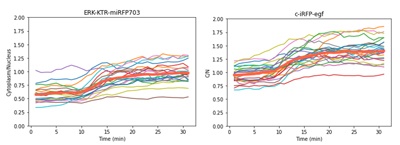
When being activated, KTR will move out of the nucleus, resulting in a decrease in nucleus intensity (Figure 2).
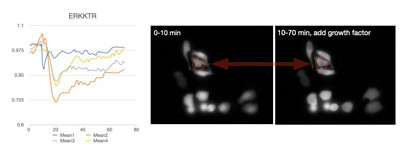
The growth factor can activate ERKKTR whose cytoplasmic relocation will make the nucleus darker. As shown in the left graph, a significant intensity drop was captured when EGF was added at 10 min.
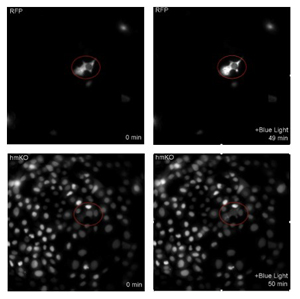
We first selected eight kinase plasmids: RAD1, MEK, KSR1, JNK1, ASK1, TAK1, p38a, and CDK1. Then I introduced those plasmids into the CRY-2 template by LR gateway reaction. The integrated plasmids will be transfected into Hela cells that have four types of KTR inside, ERK, AKT, JNK, and p38a, respectively. To visualize kinases behavior under a microscope, eight plasmids were encoded with mScarlet, a red fluorescent protein, and four types of Hela cells with mKO, an orange fluorescent protein. If MAPKK or MAPKKK can activate MAPK, the droplet will form and there will be a dark change of nucleus; if there is no relationship between two types of kinases, only cluster formation will be captured. For example, MEK1 cluster was captured and at the same time, a change of JNKKTR nucleus had been found, which identifies that MEK1 can activate JNK (Figure 3).
Apart from the scientific project, I want to say thanks to all lab members and students in Aoki sensei’s lab. Even when back to Kyushu, I still dreamt about the day we ate Nagoya fried chicken and talked until 10:30 p.m., the restaurant we enjoyed donuts together, the whiteboard had many Hokkaido maps on it, and lunch boxes that stored our short talk every day. Many memories pop up in my mind when thinking of Okazaki. I learned a lot from their attitudes and determination towards science and become firmer to be a person like them in one day. The internship here is not one sentence on my resume, but something that can push me to move forward, and I will cherish it for the rest of my life.
Message from Interns 2020

Imperial College London, UK
Sea Hee Yook
My research primarily focused on examining the proteins involved in controlling the cell cycle of Schizosaccharmyces pombe, otherwise known as fission yeast. The progression of biochemical events from G1 to M phase in its cell cycle is known to be regulated through a single CDK-cyclin complex composed of Cdc2 and Cdc13, alongside the activation of additional protein substrates at distinct timepoints.
To examine such properties, I first integrated GFP, a green fluorescence protein, at the 3’ end of a chromosomal open reading frame so that the resulting gene construct will fluoresce in varying intensities depending on the protein concentration. Confocal fluorescence imaging was subsequently conducted to examine the oscillations in fluorescence intensity over the duration of 16 hours. Moreover, I examined the expression of genes tagged with mNeonGreen, a fluorescence marker that is known to be up to five times brighter than GFP. This enabled me to visualize the expression patterns of genes such as cdc25 that fluoresce at much lower intensities.
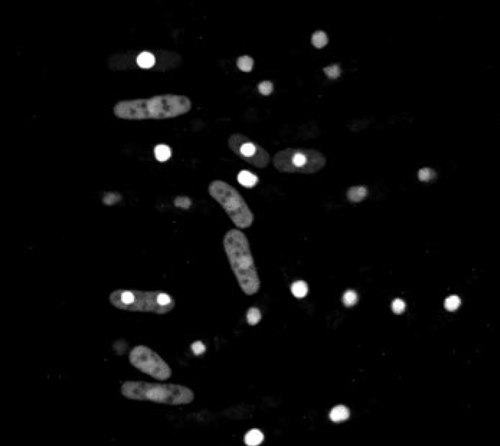
The resulting data obtained was quantified using the ImageJ software and graphed to present the time-specific changes in protein concentrations. This allowed deductions to be made on the roles and properties of individual proteins in regulating the cell cycle. The research can be further developed by attaining additional measurements such as protein synthesis and dissociation rates which are involved in an ordinary differential equation that models the S. pombe’s cell cycle. This will enable the representation of the molecular mechanisms involved in the cell cycle of S. pombe through a mathematical model with numerically quantified parameters.
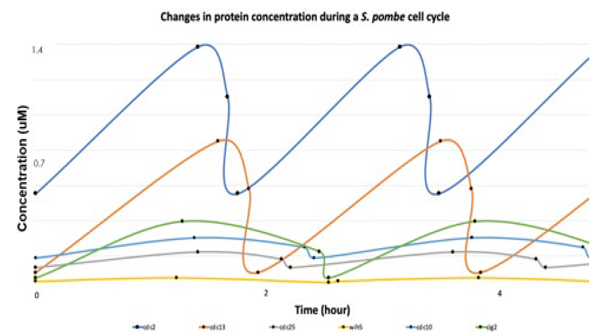
The point at which the cell begins its cell cycle, shows maximum fluorescence, and undergoes nuclear division were measured and averaged to be graphed on Excel. A decrease in concentration in the following order of proteins can be deduced from the graph: Cdc2, Cdc13, Cig2, Cdc10, Cdc25, and Wih5.
Whilst being engaged in using various imaging tools, I was also introduced to basic laboratory techniques such as western blotting, transforming plasmids into cell lines, designing PCR primers, and conducting swift but precise DNA work. Additionally, I learned about the importance of performing detailed experimental planning, careful observation, interactive discussions with other researchers, reliable data analysis, and effective troubleshooting to develop as a well-established researcher capable of presenting influential scientific discoveries.
Most importantly, I want to thank all members of Professor Aoki’s lab for allowing me to be immersed in the internship program in a warm, welcoming, and ever so supportive environment. My questions were never responded with a sigh or ignorance, but instead with detailed explanations and demonstrations.
Undoubtedly, the internship program provided me with an opportunity to further explore Japanese culture by encountering people with different values, interests, and lifestyles. Interestingly, these differences were especially evident in our lunch menus. In contrast to my lunch box which was composed of Mac n cheese, oven-grilled chicken and a clubhouse sandwich, the people at the laboratory packed their “bentos” with “yakisoba” or stir-fried noodles, “karaage” or Japanese fried chicken, and “onigiri” topped with “furikake” rice seasoning. Although there were differences in our diets, the conversations held over the lunch table allowed me to broaden my perspective on cultural diversity.
Waking up in the morning to be greeted by the sounds of birds chirping, taking a refreshing 7 minute walk from the Mishima Lodge to the laboratory, occupying myself with scientific wonders alongside other researchers, and going back to the lodge at the end of the day to try out different Japanese recipes for dinner… such routine filled my days at Okazaki with joy and excitement. I want to thank the NIBB for providing me with such invaluable opportunity, and the people at Professor Aoki’s laboratory for their unconditional support throughout the internship. I am confidently returning home with newly acquired skills and attributes that have equipped me to continue pursuing my passion in research.
Message from Interns 2019

University of Heidelberg, Germany
Bettina Welz
During my time at the NIBB in Okazaki I was introduced to new laboratory techniques and the handling of medaka in general. Moreover, I had the possibility to discuss and exchange opinions and experiences with leading researchers in the medaka research field. Besides performing a linkage mapping experiment, in which the relative position of genes to each other was analyzed based on recombination frequency, I performed cryopreservation of sperm and artificial insemination. After the dissection of medaka testis, the sperms can be stored in liquid nitrogen for long periods and used for breeding purposes. In addition, I could get insights into a germ cell transplantation technique for medaka. Further on, I got the chance to join Prof. Naruse and Assistant Prof. Ansai for a memorable fieldtrip, in which we were able to catch wild medaka within different habitats. It was such a great and unique experience for me to see medaka in their natural environment.
Besides getting new exciting scientific insights and further develop my practical skills in the lab, I got to know the fascinating Japanese culture. Experiencing the cheerful atmosphere at several summer festivals with impressive fireworks and traditional Japanese dances and music, visiting breath-taking shrines, castles and temples and falling in love with the extraordinary and tasteful food was stunning. Getting in touch with all of these amazed me more than once.
All in all, I had a great time here in Japan. Both from the research point of view, as well as from a personal perspective, I can take home a lot. I am really thankful for the opportunity to work and life in Japan and highly recommend the NIBB internship program. My personal thanks go to Prof. Naruse who welcomed me warmly in his lab and never got tired in teaching me inside or outside the lab. Moreover, I would like to thank Assistant Prof. Ansai for the inspiring discussions and his support for experimental set ups. Last but not least my sincere thanks go to the rest of the staff, especially to Naoko Torii, Ikuyo Hara, Azusa Yangawa and Toko Yamazaki for the always friendly and helpful manner. Arigatou gozaimasu!

Waseda University, Japan
Chang Yong Park
In her stem cell laboratory, I had five major experiments I was involved in : Investigating duration of cell cycle and S-phase length(with Dox and without Dox), E.coli competent cell preparation/check, Sera Lot check (ES handling), ES cloning, and Lymphocyte cloning. On the First day, I was very worried I would not be able to follow the procedures of the experiments, for I never had any lab experiences before. However, thanks to careful supervision and kind guidance of Prof. Tsubouchi and her lab members, I quickly got used to it.
Despite the shortage of time, I was really happy that I could actually get meaningful results from what I did in Prof. Tsubouchi’s lab. This was the first time I was in a big project in science and I am grateful that Prof. Tsubouchi and her lab members provided me a very pleasant first unforgettable experience in a laboratory.
The lodge I was in very pretty wide and clean, and the city Okazaki was peaceful. I did not have any major difficulties while I was there.

University of Washington, USA
Eichi Toyoizumi
I was very much intrigued by their research on visual system using deep neural network, however as I have mentioned I was a biology major with limited experience in coding. So my intention was to be exposed in the field of artificial neural network research and to translate the experiences into my future research. Due to some scheduling difficulties my internship duration was little over a week, but it was more than enough to start the research.
I was able to run/program deep learning algorithms to read the sequence of images that I created and make some observations on how they are perceived within the network. But really what amazed me the most is the fact that there is a web-based platform that enables those research to be done remotely allowing for long distance collaboration without one’s presence.
I would like to thank NIBB internship program for the support and Watanabe Lab members: Dr. Watanabe, Dr. Nishiumi and Dr. Kobayashi not only for assistance in research but for all the interesting and valuable discussions. My experience here at NIBB was very positive and will help me through my future research efforts. I look forward to our future, remote for now, collaborations.
To prospective students, I can whole heartedly endorse NIBB internship program to any of you (who are interested in life sciences of course). One piece of advise is that I encourage you to contact the faculty of your interest even if they are not listed as a lab accepting students (my lab was not listed). NIBB has great facilities that equals top research institutions in U.S. such as my alma mater, and so you are in for a treat should you be select to intern.

Vietnam National University HCMC, Vietnam
Nguyen Hoang Yen Nhi
During these two months of stay, not only have I just gained more scientific experience but also the chance to visit many places in Japan every weekend to see how beautiful and fascinating Japan is. I am very grateful to NIBB for granting me this opportunity. This invaluable research experience, once again, confirmed my determination to pursue a Ph.D. in algal biotechnology. After this trip, I have become a more mature person with strengthened knowledge and inspiration. In addition to this, I met some great friends that treated me like a family member that I would never forget. I really appreciate their kindness. NIBB internship program is a great opportunity for any research-oriented student to find the path that leads to important career opportunities. The experience and relationships I had during my stay are very precious to me and will be the step stones for me into the scientific world.

Vietnam National University - University of Science, Vietnam
Le Thu Hang
The NIBB internship program is a precious opportunity to me. Thanks for the program, I was able to experience a professional research environment as well as Japan culture. Three months spending in Japan is a great time that I will not forget. Friendship and memories come to me unexpectedly, which I bet I will miss a lot when leaving.
I was led to dig into new concepts, to broaden my scientific view in a professional and supportive environment. I would like to send my sincere thank to all members in Nakayama Lab, to my dear friends and mentors for your kind, friendly help, for giving me a warm welcome since the very first moment I came. Please let me express my gratefulness to Professor Nakayama, who has always been patient with my mistakes and taking care of me during my stay.
The internship brings me motivation and confidence to my future plan. Thank you a lot for the opportunity and I hope that in the near future, I could visit NIBB again.

Kasetsart University, Thailand
Lueacha Tabtimmai
Under the supervision of Professor Aoki, he is an open-minded person. We first shared the works that we have done so far for getting an idea. I got a chance to create my own project whilst he always shaped the idea and gave me a comment.
Without Aoki-sensei and lab members, I would not have amazing learning. I gain so many valuable experiences during staying in Aoki-lab. I am thankful for all you guys that always give me a ton of warm heart moment. We shared so many cultural things together every lunchtime. Besides from hard-working culture, there are so many activities here that make me really enjoy every day. Onada-san, is a lab secretary who always gives me a hand when I got in trouble. I really appreciate all help from everyone that never denies any inquires. Overall, NIBB gives me the best opportunity for being a part of Aoki lab. I definitely recommend NIBB program for all students who would like to get a good experience in Japan.

Mahidol University, Thailand
Neen Phan-udom
I was under the supervision of Asst. Prof. Ebine during my internship. My work mainly consisted of performing transient assays for Arabidopsis thaliana leaves and visualizing the localization of the targeted proteins via confocal microscopy. I learned to find and take pictures of the cells expressing GFP/RFP-tagged proteins to later measure their activities and possible interactions. I also got to attend a 2-day summer practical course on oil bodies in liverwort which included mini fieldwork to collect liverwort samples and visualize their oil bodies under the microscope.
In addition to gaining more knowledge and laboratory skills, this internship was one of the most wonderful experiences I’ve ever had. I received a very warm welcome from all the lab members as well as other staff at NIBB. I was pleasantly surprised when the lab organized a welcoming party for me and my lab partner on our first day (which has gotten me really into soumen). Everyone was so nice and friendly which made me feel very welcomed and look forward to my tasks every day. I was on the verge of crying on my last day here when I had to say goodbye. I also had many wonderful moments learning about Japanese culture both inside and outside of the lab, not to mention having the privilege to live in Okazaki city and do some traveling on the weekends. Japan really is a beautiful country.
I will never forget my two weeks here. If anything, I wish I could have stayed longer and learned more about membrane trafficking in plants. The internship has shown me the beauty of basic research of exploring the unknowns and left me wanting for more. Really, I can’t thank everyone enough for this amazing experience. I am forever grateful for this internship opportunity.

Kyushu University, Japan
Nguyen Quang Khai
The laboratory that I joined is the Stem Cell Laboratory. Under the instruction of professor Tsubouchi, I participated in two on-going pieces of research: Embryonic stem cell S-phase length measurement and dNTP-recognize complementary protein development. From the experiment, I received many opportunities to practice basic cellular techniques, such as complement protein, cell cultivation, colony identification, etc., that I learned previously.
Another memorable aspect of the internship was experiencing research student life. Every day, I went to the lab at a set time. Initially, professor Tsubouchi described the tasks we tried to accomplish on that day and lectured about the ground understanding of the subject matter, then we proceeded with the job, and after we finished all the experiment on that day, we had small seminars with other lab members. During the small lectures and the seminars, I felt intrigued by the discussion, questions, and answers, and these were the times when I studied new things about stem cells. On some occasions, I needed to be in the lab at night to prepare for the long, time-sensitive experiment. On certain days, the task was only analyzing live-imaging videos of cells, which is far more demanding than it sounds. I must say the life of a researcher is not an easy one. It is difficult and arduous, yet, the moment when I received the expected result, it was very rewarding and worth the effort.
From this experience, I have learned many new things. I have studied the technical knowledge, practiced the skills related to cellular biology, experienced the life of a researcher as I mentioned. More than that, I consider my professor to be a role model of a researcher, being someone who is well organized, dedicate, understands the study thoroughly, and passionate about science. After the internship, I can confidently say that I want to pursue an academic career, and I believe that I can do so.
I want to express my gratitude to professor Tsubouchi and other lab members, the staff members of the dormitory and the institute. Everyone contributes to my highly enjoyable experience at NIBB.

Kyushu University, Japan
Penpatchara Silaguntsuti
This internship was an invaluable and memorable experience. I learned and practiced in several experiments on protein detection using fluorescence microscope and immunoblotting. It was the first time to observe my own plant protoplast samples and scanning with fluorescence microscope under supervision of experts.
Since the first day of arrival, I received a warm welcome from everyone in the laboratory including the supportive staff. It was my honoured to attend Professor Ueda’s lecture on current research focus—genes that responsible for Golgi network transportation in Arabidopsis and oil bodies in liverwort. Afterwards, I was surprised about welcome party and hospitality in the conversation.
I learned a lot of experimental skills from the very essential thing like keeping the protoplast alive, protein extraction, SDS-PAGE, immunoblotting and how to use fluorescence microscope. More than that, the first hand experience from being trained by Assistant Professors of the laboratory was something I could not find from somewhere else. I would like to give special gratitude to Professor Ueda who accepted my application and Assistant Professor Ebine who was my supervisor to most of my experiment and was skillful at troubleshooting the problems. I felt thankful to meet Assistant Professor Kanazawa for his summer school program, Mrs. Okubo for her kindness as a secretary of the laboratory, Mr. Minamino who possess fluent English skill and humour, Mr. Hatchinoda and Mr. Norizuki for his hospitality. I also wanted to thank other few laboratory members that I hardly talk to them for their kind gestures. Last but not least, thank to Mr. Glen, the supporter staff, I received guidance about NIBB and some advice how to adapt myself to Japan.

Szent Istvan University, Hungary
Sarah Shaqiri
The Shigenobu Sensei’s lab has been studied the symbiosis between aphid and the endosymbiont Buchnera aphidicola. They’ve published several papers on transcriptome of aphid Buchnera and being part of it for a period of almost two months I had to compare the transcriptome between different aphid morphs. It was quite interesting topic yet challenging experience for me because previously I worked on plants and there I was going to work on aphids particularly doing the Next Generation Sequencing for the first time. The project included both computational analysis (bioinformatics) as well as molecular work (bench work).
During the molecular work I was working together with Dr. Nozaki a post doc of the lab. Here I was able to conduct a range molecular experiments like RNA extraction, RNA quality evaluation, RNA library preparation, Real-Time Quantitative Reverse Transcription PCR, tagmentation and generating sequence library using Nextera XT DNA library kit [Illumina], from which we successfully passed this part of experiment. Heading further into computational analysis I faced difficulties but I would want to thank Shunta Yorimoto PhD Fellowship in our lab which helped me giving the best explanations so that I can work further on in bioinformatics in life. I also want to express my graditute to Shigenobu sensei which guided me to be familiar with Unix commands, R commands, DGE-analysis and more. Despite the fact that Sensei had a busy schedule he always found time to help me and give advices to me.
Taken together I had a wonderful experience at the NIBB and in Japan in general. Apart from being welcomed from people in the lab I also made new friends and adjusted myself in the whole different and unique culture of Japan. I am glad I came to Japan and I believe there is much more from it that I could explore. In the end I hope a lot of scientists would have the opportunity to gain experiences like this in the future. It was worth every second.

University of Hawaii at Manoa, USA
Vânia Filipa Lima Fernandes
Over the period of two weeks at NIBB, I was able to enhance my knowledge regarding histological techniques complemented by software analysis that will be used to reconstruct the Astyanax brain in 3D.
Professor Fujimori and Oka Sanae’s expertise and experience alongside with everyone in the lab were crucial for the progress of this project. All lab members were very receptive and helpful which created a very friendly and inviting but also prolific atmosphere. This experience contributed immensely to my project and my personal growth as a scientist, making me more motivated to learn and explore other perspectives.
Overall, NIBB Internship allowed me to experience a new scientific environment and open my mind to future perspectives in Japan and I will always be grateful for the amazing opportunity.
Message from Interns 2018

University of Sciences and Technology of Hanoi
Nguyen Ha Ngoc Anh
My research revolves around lutein production in microalgae. Lutein is a valuable biocompound, playing a major role in eye protection and hold a promising future in the food supplement industry. This is a long on-going research and I took multiple approaches towards boosting lutein accumulation in Chlorella sorokiniana, regarding from strain selection, media optimization to mechanism verification.
Despite the lack of time, this journey has plenty take away lessons that I hold at heart. First and foremost, I would like to express my gratitude to Minagawa-sensei, my mentor Satou-san and all lab members for a wonderful 3 months. Not only did they welcomed me with the warmest hearts, they also introduced me to a nurturing science community, where lab members can deliberately discuss and groom each other to success. Even though I am still young and rather inexperienced, all lab members always come to my aid, give me advices to help me freely test out my vision, provide me the opportunity to take multiple approaches in the matter. Furthermore, during this trip, I was able to make invaluable friendship with people from all around the world, who all had inspired me in every way possible. The spring of 2019, filled with sakura, pot-luck parties and laughter will always be the memory that I treasure.
In conclusion, the NIBB Internship is a remarkable, high-quality opportunity for all biology students who wishes to embark on a scientific career. I would like to say thank you to everyone at NIBB who had made this journey possible for me and I wish everyone the best of luck in the future.

University of Santo Tomas, Philippines
Sheena Josol
In my 2 months stay, I was given two research projects which independently aimed to (1) reconstitute tumorigenesis and oncogene addiction in a manner dependent to Trimethoprim and (2) dissect molecular mechanisms of stochastic ERK activation, both of which involved the use of wildtype MCF10A cell lines. In the first project, the oncogene of interest that we intended to induce are the ones involved in ERK signaling pathway:the G protein KRas and the protein kinase BRaf specifically its KRasG12V and BRafV600E mutant version respectively which are widely implicated in several human cancers.
The first step in inducing oncogene addiction was to establish the needed cell lines which in this case is a Dihydrofolate reductase (DHFR)-expressing MCF10A cell line. As we all know, the best way to ascertain the function of one biomolecule is to remove it from the system and see what happens next. The TMP-DHFR system makes it all possible. This model system works in such a way that it requires the protein of interest to be fused to a destabilizing domain like DHFR which targets it for degradation. The protein is then rescued by the addition of TMP that binds the DHFR and inactivates it which in turn enables it to regulate the stability of protein in a rapid, reversible and tunable manner.
Through imaging, western blotting and cell proliferation assay which I spontaneously did during the course of my internship, I was able to analyze the stabilizing effect induced by TMP, quantify the expression levels of induced oncogenes and examine whether our selected cell line acquire cancer like phenotypes after oncogene induction respectively. As for my second project, due to limited time, I was not able to achieve the main goal of studying the stochastic ERK activations.
I am beyond grateful to every member of the lab, who humbly imparted their knowledge to me especially to Aoki-sensei and to my mentor Reina-san who patiently taught and guided me with everything. I would also like to express my gratitude to Onoda-san, the secretary in our lab, whose unfailing efforts made my first ever Autumn experience the best. I really appreciate the good laugh I’ve shared with everyone during lunch time and all the parties they have prepared just for me.
Overall, this experience has not only filled me with significant learnings in the field of science but also let me gain great memories, learn meaningful life wisdom, establish lasting friendship with people from different parts of the world and lastly brought me to wonderful places that made me appreciate the beauty of Japan more. Indeed, Japan is a beautiful country but more than everything else, I think the culture of hardworking, disciplined and kind people makes it more special.

Mahidol University, Thailand
Watcharin Unwet
I had a wonderful time during internship program period. In 15 days, I learned a lot of skills including image analysis using FIJI (A distribution of ImageJ software program which includes many useful plugins contributed by the community) which is a very useful tool for biologist and learned some essential laboratory skills for plant biology such as molecular biology technique, using confocal microscopy to study cellular morphology of plant, using Katikati2 software program for counting the plant epidermal cell and palisade cell and so on. These techniques can be applied to my senior research project in my university.
As a biologist who loves to study plant science, I felt really enjoy working in this lab. I would like to thank all Kawade’s laboratory members: Kawade, Tomoi, Nozaki, and Yamaguchi for everything. I also enjoy my daily life in Japan, I had learned some basic Japanese sentences and their cultures. Moreover, I had met with so many foreign friends there and had a great time together.
I would like to give a big thank to NIBB internship program for making this memorable experiences possible and I would like to encourage those students who interested to study biological science and learn some Japanese culture. For me, this internship program is one of the most valuable moment ever.

Kyushu University, Japan
Tsukasa Ryu
Medaka is a small freshwater teleost fish that are now developed as a model organism for vertebrate. Given that its genome sequence is now available on database, medaka is ideal for genetics research since they lay eggs in very frequent pace. It is my unforgettable memory to see a breeding sector full of different strains and generations of healthy medaka. Variations in phenotype of different medaka strains were very interesting to compare.
During this week visit in Prof. Naruse’s laboratory, I experienced a set of experiments that aimed to observe two body color mutant phenotypes of medaka to understand genetic linkage and polymorphism by measuring genetic distance based on loci of certain genetic markers. The set of experiment included the sorting of F2 embryos, fin clipping of F1 parents, genomic DNA extraction, PCR and restriction enzyme treatment of PCR product. In addition to these experiments, I also had a chance to experience a brief training on microinjection technique using CRISPR-cas9 method. These sequence of experiments taught me the logics of constructing a convincing analysis.
Moreover, the program was not limited to the experiment. Additional lectures on background knowledge of the experiment, efficient use of database, and on primer designing method are some of the things I learned during this internship. Those practical knowledge I gained from the lectures are now supporting me in my laboratory.
In this extremely short period of visit, I was able to learn countless amount of knowledge from very packed content of the internship. I really appreciate Prof. Naruse, Assistant Prof. Ansai, all laboratory members, and internship staffs for preparing such a wonderful program. It is my honor to say that I have completed an internship at NIBB, especially in this laboratory. Even from this short visit, I can proudly say that I have learned so many valuable things from every experience I had during the internship.

Kyushu University, Japan
Do Thuy Linh
From the document provided by Shigenobu sensei and guided by Mr. Shunta Yorimoto, I have learned about UNIX commands and BLAST. After successfully identification of BCR3 and LSZ-i homologs in C.japonica transcriptome, with the enthusiastic and conscientious support from Dr. Chen-yo, who is doing Postdoctoral Fellowships in our lab, I was able to conduct a wide range of molecular experiments from RNA extraction, RNA quality evaluation, Real-Time Quantitative Reverse Transcription PCR, in situ hybridization, using the mRNA transcriptome sequence of bacteriocytes. The result indicated that both BCR3 and LSZ-i show the gene conservation in C.japonica.
There were a lot of other things about this program that were worth being excited about. First of all, the sheer idea of spending my summer in NIBB thrilled me: it’s the centre of research, of state of art technologies, of professional experts and of friendly people. Shigenobu sensei is so supportive and took care of me very much. Although my professor is really busy with so many business trips every week, he still asked me: "Apart from science, would you like to visit some sightseeings in Okazaki?". Then he and his wife took me and Mr. Chenyo to go visit both traditional and modern sightseeing in Nagoya, and also drink coffee and have many talks about life and science careers. Moreover, I also met a lot of interesting people in my program: Miyuzu-san, Bino-san, Ichikawa san and other staffs in the Common Facilities. They were and are still my inspiration.
After this program, I feel contented that I have got the chance to get to know scientific research in many aspects. Each of the techniques and knowledge acquired in the program will serve as a door to an educational pathway that I may choose to pursue later on during my study at Kyushu University or beyond. In the end, I felt so honored to be selected for the internship in NIBB and Assoc. Prof. Shigenobu Shuji’s lab and I was in the best research environment ever.

Peking University, China
Peng Chen
During my stay, I did some researches on auxin biosynthesis in the moss Physcomitrella patens (Physcomitrella). The experimental material was a mutant, showing defects in periclinal cell divisions, which is one of the most significant developmental process for land plants.
I must express the depth of my gratitude to Professor Mitsuyasu Hasebe for giving me the opportunity to learn and work in his lab. He is an erudite scholar and a kind supervisor for students. And I should show my appreciation to Dr. Tsuyoshi Aoyama who guided my researches in those two months. He answered every question I asked patiently and taught me a lot about Physcomitrella and its experiments. Also, many thanks to all the members in Professor Hasebe’s Lab. You were glad to give me a favor every day and showed me an outstanding, efficient and optimistic scientific research team. Specially, I would like to say thank you to Mr. Cowan in international cooperation office, for his responsible attitude towards all the procedures during my intern and his concerns for my daily life.
This was my first time being aboard, but I felt at home in Okazaki. I love this quiet and peaceful city and I think it is one of the best places for scholars to study their scientific topics. I spent an unforgettable time in NIBB and I hope that I can continue my graduate education in Professor Hasebe’s Lab.

Atma Jaya Yogyakarta University, Indonesia
Astrid Valerie Putri Kusuma
I worked on the project for characterization of HP-1 like protein in Tetrahymena. HP1 is a conserved chromosomal protein that forming higher order of chromatin structures and binds to H3k9me (lysine 9-methylated histone H3). HP1 proteins have an amino terminal CD and a carboxyl-terminal CSD. There are more than 20 known heterochromatin protein but we still do not know how they interact with each other.
Therefore, the tasks that was given to me is to construct plasmids for expressing 12 CDs in Tetrahymena HP-1 like proteins. These will be used to obtain 12 CDs in Tetrahymena HP-1 like proteins and the final goal is to analyze chromodomain’s affinity for H3K9me3 or H3K27me3 by using isothermal titration calorimetry.
My internship so far is extraordinary. I am truly honored and blessed to work under Nakayama Sensei. He is such a great person and always willing to help and assist me in doing my project. This kind of humble treatment is one in a million and is not famous in my country. The other staff members are also really kind. Thanks a lot to Ken, Aki, Asai, Yuriko, and Kyoko for being supportive and patient with me. All of them were really helpful in helping me working in the lab and for any other administration.
I also feel grateful for my accomodations. Although my arrival was planned in such a short time, but they manage to deal with all of it. Special thanks to Glen who took care with the administrations regarding my arrival and accomodations. The people here are so nice and it’s really easy to make friends.
Last but not least, this is one of the great experience that I will always remember. Though it sounds exaggerating but when treatment with kindness is almost in every part of the process, this will make someone always remember. I hope that I will be given the chance to be back here someday. Terima kasih banyak !
Message from Interns 2017

Kyushu University, Japan
Thi Tran Ngoc Minh
ESCs have a unique cell cycle of shortened gap phases compared to other cell types. Analyzing the differences in DNA replication (S phase) and mitosis (M phase) of ESCs and differentiated cells prove a useful approach to elucidate how and why ESCs proliferate with such unique cell cycle. In one experiment, I assisted in sorting ESCs and differentiated fibroblast cells at different S phases, from which samples will be used for further analysis in their replication process.
Despite the short time, thanks to the kind instruction of all the lab members, I was able to learn a wide scope of laboratory techniques such as cell sorting, FACS analysis, confocal microscopy, live imaging analysis, tissue culture handling of various cell lines.
I would like to express my greatest gratitude for Tsubouchi-sensei and all the members of Laboratory of Stem Cell Biology for their very kind guidance, delightful conversations on Kansai culture, scientist life stories and the wonderful misokatsu cuisine. Apart of lab work, the wonderful accommodation arrangement at Mishima lodge has made my stay very comfortable and helped me make unforgettable friends.
At last, I am eternally grateful for NIBB to give me the opportunities to work alongside with such hardworking, kind and talented scientists in the Laboratory of Stem Cell Biology. The research atmosphere here is truly inspiring and exciting and I really hope to come back here in the future.
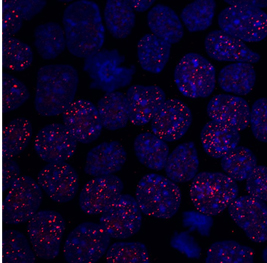
Figure 1 DNA damage tracking by observing Rad51 (red) and chromosome (blue)

Pontificia Universidad Javeriana, Clombia
Andrés Julián Gutiérrez-Escobar
1- MBGD: orthologous detection.
MBGD is a database system developed to compare different features of bacterial genomes, such as: gene order, motif identification and orthologous/paralogous detection. I have received training about how to use not only MBGD but also the RECOG tool to extract the core genome orthologous alignment from 141 H. pylori genomes.
2- SNP calling and the vcf format
The full genome and the concatenated core genome sequences were used to create vcf files for each strain using the snippy tool. Finally, the vcf files were merged independently using vcftools.
3- Imputation and fineStructure analysis
Each merged vcf file was imputed using BEAGLE and the fineStructure software was used to obtain the ancestry matrix for this subpopulation.
4- Positive selection analysis
Finally a whole genome positive selection analysis was conducted using PhaME.
5- Scripting
Professor Uchiyama teaches me how to write scripts in order to automate all the analysis using bash shell.
About my experience in Japan
Japan is a beautiful country full of kind, humble and educated people. I am just amazed about how good is this precious country. I was hosted at the Michima lodge. This place is cozy, silent and friendly. I have felt like in home.
My internship was directed by Uchiyama sensei. He is not only one of the best bioinformatician in the world, but also one of the best human beings I have ever met. From him, I have learned extensively not only about genomic but also how to be a better person. I also want to thanks the Office of International Cooperation during and after the internship program. In general, the NIBB is the best academic place I've ever known in my entire life. Here, you can learn how to do the best science. I hope I can return one day to this incredible institute.

Mulawarman University, Indonesia
Anisa Fitri Rahayu
In fission yeast, such as S. pombe, there are two HP1 family proteins (Swi6 and Chp2) and play as main factor for maintenance gene silencing at heterochromatin regions. These HP1 proteins recruit other binding partners in order to maintain the repressive structure of heterochromatin. The exact means by which such HP1 family proteins recruit distinct set of partners, however, remain incompletely understood. My study in this internship program consisted of disturbing the function of one of Swi6-interacting proteins and characterizing mutant strains by silencing assay. The effect on heterochromatin integrity and stabilization can be check by growing the cells in medium containing low level of adenine, because on this study I used the S. pombe strain carrying ade6+ marker gene in the heterochromatin region on centromere 1. The isolated cells showed variegated phenotype on the medium resulting in white and pink colonies and indicated that silent chromatin at centromere is less stable in the mutant strains.
I feel lucky I got the opportunity to join the program because I got some precious experiences not only about lab work but also about Japanese culture. I had chance to attend the laboratories join meeting such as paper and progress research presentation also watched seminar from other laboratories. All the knowledge I got there was new and could answer my curiosity about heterochromatin also made me understand some concepts that I had not understood before. I hope the experience and knowledge I got from the program can help me to pursue my career plan as researcher in the future. I got nice atmosphere too in the lab and I really enjoy my time there, I would like to thank for all laboratory members: Machika, Naoko, Kyoko, Yuriko and Ken for helped me in the lab and being friendly, especially for Nakayama sensei for the guidance in lab work also helped me understand all about epigenetics, heterochromatin as well as helped me to prepare my entrance examination to enter NIBB graduate program (SOKENDAI). I hope to have opportunity in near future to come back and study at NIBB.

Heidelberg University, Germany
Bea Riebesehl
Already during my bachelor thesis in Heidelberg, I worked with the Japanese rice-fish also known as medaka. Proven as a highly suitable model for transgenesis experiments, I wanted to expand my knowledge about this sweet water fish and improve my experimental techniques in the laboratory. Therefore, I applied for an internship in the laboratory of Prof. Kiyoshi Naruse studying the genetics underlying pigment cell and neural development in medaka.
During my 2-months stay at the NIBB in Okazaki, I focused on the development of a moto neuron reporter line by using CRISPR-Cas9 mediated knock-in of a green fluorescent protein. Therefore, I designed sgRNAs targeting the 5’UTR of neural genes that are exclusively expressed in the brain. These sgRNAs were conjected with Cas9 mRNA, a donor plasmid containing a Tbait sequence and a Tbait sgRNA to open the donor vector. Unfortunately, the chosen genes seemed to have an essential role in embryo development, so most of the injected embryos did not survive until hatching. I optimized the system by targeting more sequences 600-200bp upstream of the transcription start side and by adjusting donor plasmid and sgRNA concentrations. Luckily, promising neuronal GFP expression could be found at last. Additionally, I used a donor vector with a DsRed flanked by LoxP sites followed by GFP. Upon Cre recombination i.e. by crossing to a Cre expressing line, DsRed will be excised leading to GFP expression. The injected fish with mosaic fluorophore insertion are now growing and will be tested for exclusive moto neuron expression in the next generation. Eventually, they will become founder of an inducible moto neuron transporter line.
Facing obstacles, I was obliged to think about how to design and conduct experiments to prove hypotheses and optimize the experimental procedure. Working independently on my project helped me to gain self- confidence and comprehension of scientific research. Hence, I am very grateful that Prof. Kiyoshi Naruse gave me the opportunity to work in his laboratory. I would also like to thank Satoshi Ansai who was always there to answer my questions and all the lab members who helped me to find my way in the lab. Special thanks to Chieko-san for explaining me about Japanese language and culture during lunch breaks. Finally, I would like to thank the NIBB internship program for its financial support.
My stay in Okazaki was both enriching and inspiring thanks to all the people I met on the way. Coming to Asia for the first time in my life, certainly changed my thinking and understanding. I am very glad that I came to Japan and hope that a lot of students will benefit from this program in the future.

Universiti Sains Malaysia, Malaysia
Kathrine Tan Xin Yee
I involved in the project entitled ‘ Identification and Quantification of Symbiotic Bacteria in a Social Aphid, Ceratovacuna japonica by metagenomic approach’. C. japonica is a species of eusocial aphid. One of the distinctive features of this aphid is the evolvement of soldier caste in the colony. This unique feature has distinguished C. japonica from the model species, Acyrthosiphon pisum and it is possible that the symbiotic bacteria presence in the soldier caste and reproductive caste of C. japonica as well as the A. pisum are different.
Throughout the three weeks of internship, I tried to identify and quantify the secondary symbionts that present in the reproductive caste of C. japonica. The first week of my internship was spent in conducting laboratory works such as extracting genomic DNA from the reproductive caste of C. japonica. Preparation of 16S metagenomic sequencing library was conducted in the second week followed by sequencing of metagenomic library using Illumina Miseq sequencer. I would like to thank Ms. Miyuzu Suzuki and Ms. Asaka Akita for assisting me in the experiment. While waiting for the results of sequencing, I spent most of my time in preparing the workflow for the analysis of metagenomics data. As I am not familiar with writing scripts and UNIX commands, therefore I spent quite a considerable time in learning how to operate data analysis pipelines like QIIME. However, with the help of my dearest lab member, Mr. Shunta Yorimoto, I was able to perform simple data analysis and certain data processing commands. I am glad that the analysis demonstrated the possibility of finding novel symbiotic bacteria in C. japonica.
Apart from that, I am thankful to the members of NIBB Core Research Facilities, Functional Genomics Facility Department. Without them, I do not think that I could adapt to this new environment in such a short time.
Terima kasih and ありがとうございました。
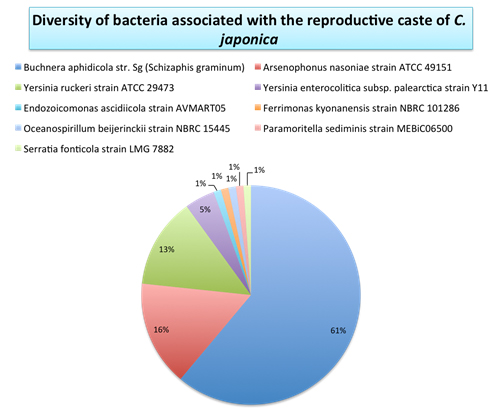

Heidelberg University, Germany
Kerim Anlas
Prof. Naruse’s laboratory of Bioresources provides access to numerous medaka (Oryzias latipes) strains as well as a plethora of other Oryzias subspecies. While closely related, these fish have, for instance, not only exhibited disparate mechanisms of sex determination but also distinct pigment cell (chromatophore) type occurrence. Thus, they constitute suitable model systems for studying evolutionary processes.
Whereas higher mammals feature only one chromatophore type (melanocytes), several are found in teleosts (e.g. black melanophores, yellow xanthophores, blue cyanophores). Another variant, the orange-white leucophores, are present only in a few fish species. They are found in medaka, however not in O. woworae, an indonesian relative. Yet, intriguingly, the genes underlying leucophore development and specification in O. latipes are conserved in the latter. This hence raises the question of how these genes have functionally evolved in O. woworae.
To tackle the aforementioned conundrum, we employed the CRISPR/Cas9 system to knock-out and label two of those genes, pax7a and slc2a15b, in both species. Pax7a is of particular interest, as it is also crucial for xanthophore development in medaka. Ultimately, several transgenic O. woworae and O. latipes lines were generated for each of the genes listed above. These either simply lack functional target genes or harbour a GFP coarsely inserted into the endogenous loci, resulting in a disruption of their respective ORF, i.e. a knock-out via fluorescent reporter knock-in, which, in turn, further enables the study of target protein localization and gene expression dynamics through life-imaging.
As to my everyday life outside of science, everything provided by the administrative office was very convenient and I felt comfortable living in Okazaki. On the weekends, I was able to explore other cities and parts of Japan which also left me with many fun and memorable impressions.
Taken together, I had a wonderful, instructive experience at the NIBB and in Japan in general. Accordingly, I highly recommend the NIBB internship program. Every member of the research group (and all other staff I met) were extremely welcoming and friendly. Hence I would like to thank each one, as well as specifically Dr. Satoshi Ansai and those who assisted me during my project. I would further like to express my gratitude to Prof. Naruse for being a very kind and hospitable host who generously invited me to join a national scientific meeting and warmly introduced me to several aspects of Japanese culture.

University of Belgrade, Serbia
Olivera Valentirović
The research I performed during my internship at NIBB was on the ciliate Tetrahymena thermophila. This was my first time working in a laboratory as well as with Tetrahymena. I found its life cycle and the extreme processes happening on its genome during its sexual reproduction process called conjugation extremely fascinating. Those peculiarities make it a really favorable and attractive model organism.
My project was to characterize the proteins involved in the formation of heterochromatin and DNA elimination during genome rearrangement within the macronucleus. In one experiment I generated several transgenic cell lines that express proteins tagged with green fluorescent protein to analyze localizations of heterochromatin protein candidates. It was a great experience seeing gene gun transformation for the first time, and then learning how to use it myself. I also learned a lot about culturing, storing and transferring Tetrahymena and how to perform a selection of the transformed cells and phenotype assortments using different concentrations of selective substances in the medium. In a parallel experiment, I aimed to reveal the sufficiency of few proteins for DNA elimination by a tethering assay. I faced some difficulties during this experiment, but I found it a good experience for my future work, because that I may face and need to solve on a daily basis. Every day I was presented with interesting challenges and new chances to find a way to untangle some problem. In the end, I managed to go through all the intended steps and successfully finish the experiments with some of the samples. I wish I could have had a longer period of time to do more experiments with different conditions, and finish all of the initially planned experiments.
Accommodation at Myodaiji Lodge was extremely comfortable which made my stay very pleasant. I didn’t lack anything and everything was professionally organized, thanks to the International Corporation Office.
All of the lab members of the Division of Chromatin Regulation were always willing to help me with my work, as well as with my stay. They gave me a lot of precious advices for which I am immensely grateful. It is a great honor for me to have worked with such polite, kind and hard-working people. I would especially like to thank prof. Dr. Nakayama who gave me this opportunity to join his lab and offered me an unforgettable experience of Japanese culture, its delicious cuisine and its people. I am eternally thankful to Dr. Kataoka from whom I’ve learned many valuable things, who patiently and carefully watched and guided me through every step and who showed me how wonderful and incredibly fun working in the laboratory can be. I really appreciate friendly and welcoming atmosphere during my stay.
This was a life-changing adventure for me. In addition to the exciting bench work in the actual lab, I’ve gained many unforgettable memories, visited amazing places, felt one unique culture, met remarkable people and made lasting friendships. The whole experience inspired me to pursue my career in science even more than before.
Message from Interns 2016

Middle East Technical University, Turkey
Emre M. Ipekoglu
I believe that I soaked up the atmosphere not only for academic aspects, but also for cultural domain which was another important gain for my intellectual development. Also, I tried to introduce my culture, it was a perfect sharing occasion, thanks to this friendly and supporting environment in the laboratory.
The organisation of the internship was very professional thanks to the international cooperation office. Accommodation in the Myodaiji Lodge was comfortable and neat, everything was well-organised in order to meet the every needs of the visitors.
I would like to acknowledge Minagawa-sensei and dear members of his laboratory.

VNU University of Science, Viet nam
Tran Thi Hong Nguyen
In my research, I planned to visualize Wnt proteins by the addition of fluorescent tags because it is known to be quite difficult to generate anti-Wnt antibodies available for immunohistochemistry. Prior to visualization of tagged-Wnt proteins in embryos, in the beginning, I focused on the optimization of linker length to minimize the effect of tags on the activities of Wnt proteins. According to some recent research, it was shown that activities of Wnt proteins are frequently damaged by the addition of fluorescent tags. Therefore, I tried to optimize the design of tagged proteins by changing the length of the liker peptide connecting Wnt to EGFP tag. Specifically, I generated 5 constructs in which the lengths varied from 9 to 55 amino acids. These constructs were expressed in culture cells and in Xenopus embryos and examined to what extent Wnt3a activity and EGFP fluorescence were retained.
During my three months of stay in Okazaki, I had lots of unforgettable memories with labmates and other friends here. I was able to learn some interesting research topics in my internship under the valuable guidance and inspiration of Prof. Takada, my mentors – Ritsuko san, Mii san, and the warm hearts of Nobata-san, Utsumi-san, Takashiro-san and other labmates who encouraged me to successfully complete my internship here. During my first visit in Japan, my flight was delayed by 2 hours and I arrived in Nagoya at midnight. It was so touching that Prof.Takada and his wife came to receive me from the airport so late at night.
Furthermore, I can never forget to mention about my stay in Okazaki would never be meaningful without the taking care of my kind-hearted host family. I would like to express my sincere thanks to Fukada-san, Makiko-san and my host brother - Shortaro-kun who always take care about my needs, be with me in many short trips to explore Okazaki, Nagoya… Thank you so much for treating me as your daughter!
Last but not least, many thanks go to my beloved parents, my “someone” for unconditional love and continuous supports me when I went abroad.
The NIBB internship program is really meaningful for international students who want an experience with one of the highest reputations in education as well as the Japanese culture. Thanks all, for everything I experienced here!

Heidelberg University, Germany
Sevinç Gücüm
There are several advantages of this internship for my academic career. First of all, O. hubbsi was a new model organism for me to apply genetics and molecular biology methods. Also, sex determination field was a new concept for me to study. In this 3-month internship we were trying to identify a new gene which determines either male or female sex in O. hubbsi. For this purpose I have done genotyping as well as histological studies. Finally, I have applied CRISPR/Cas9 gene knock-out system for the first time by myself with the help of my supervisor. This helped me a lot to understand the method and its applications.
Apart from the experiments, lab members and the administrative office was always very helpful and friendly to me. The very first week, NINS Office set a meeting for foreigners to inform us about the life in Japan and official works needed to be done by us. I found this meeting very informative and helpful. In this meeting, I have also met international students from different labs and we became very good friends. Later, we had chance to hang out all together and discover Okazaki. In my opinion, there might be even more meetings for foreign students to help them find friends during their stay. During my stay, several conferences, parties, and events took place and I have found this very helpful to get to know Japanese culture. Thanks to the localization of Okazaki in Japan, I travelled to Tokyo and Osaka to learn about the Japanese history, people, culture, and especially cuisine.
Overall, my lab work helped me to learn a new concept, apply new methods, and gain self-confidence, and more. I wish, I could even stay longer if I did not have to turn back to Germany for my thesis project. I need to thank to Prof. Dr. Kiyoshi Naruse, Dr. Yusuke Takehana, Dr. Saori Yokoi, Ms. Akiko Nishimura, and all other technicians and responsible people of the program. I am very grateful to them for giving me this opportunity to be in Japan. I hope, I can come back to Japan in the future, as well.

University of Delhi, India
Preeti Khandelwal
The topic for my internship project was “ CRISPR/ Cas mediated gene knockout to generate see-through medaka”. To generate the knockouts we used the CRISPR/Cas based RNA-guided endonuclease in cab strain of medaka. Five causative genes for pigmentation in medaka were selected namely pnp4b, dnajb14, bnc2, pisp1 and snapc3. First step is to prepare cas9 nuclease and engineered single-guide RNAs (sgRNAs). Capped RNA encoding for Cas9 nuclease is transcribed from an expression vector pCS2+hSPCas9, constructed for fish. Specific sgRNA for each above mentioned genes are engineered by cloning a pair of custom-ordered oligonucleotides in vector pDR274 and then in vitro transcribed using T7 RNA polymerase. I successfully completed the key step of my project.
Every morning freshly released fertilized eggs are collected and microinjected the one-celled eggs with the Cas9 nuclease (200ng/µl) and each sg RNA (50ng/µl) mix, using a thin glass needle held in a micromanipulator. The mRNA used is very important in deciding the role in pigmentation in medaka. After incubation of fertilized eggs at 28 ℃ for 3-5 days, genomic DNA is extracted from each egg. The target sequence are amplified by PCR using specific primers for each gene. PCR amplicons are applied to gel electrophoresis on the latest equipment, MUltiNA, which made the gel electrophoresis very simple, fast and easy to analyse. The electrophoretic results are analysed to estimate the targeted genome modifications and identification of mutants by Heteroduplex mobility assay (HMA). In case of discrimination among wild type heterozygous and homozygous, wild type and homozygous mutants show a single band with small difference in their mobility whereas heterozygous mutants exhibits multiple bands. Out of five sg RNAs , four sgRNAs induced mutations. I really wished to perform further experiments to get the end results of my project but I had only one month which was very short duration to carry out all the experiments. But everyday was a learning day for me. As a fish endocrinology background these experiments were absolutely new to me but I am really grateful to assistant prof. Yusuke Takehana, who really helped me at every step of my experiments and taught me to perform microinjection easily in the medaka eggs, which I was finding very difficult initially. He also provided me the bicycle to commute to the lab from the lodge and to explore the Okazaki. Cycling everyday helped me to stay healthy. I truly appreciate the interactive and friendly atmosphere in NIBB and prof. Naruse’s Lab. All the lab members and staff are very helpful and outstandingly supportive. I enjoyed my stay at Myodaiji lodge which is very well equipped, organised and comfortable. Its location is so apt which made my transition very smooth. I also got the chance to visit some beautiful tourist places in such as Takeshima island, Okazaki castle, Arashiyama and kiyomizudera temple in Kyoto to understand the Japanese culture. I enjoyed every day in this incredibly beautiful, safe and fascinating country. This internship aid in my overall scientific development and also from cooking my own vegetarian food to exploring rich cultural values of Japan.
I hope to have another opportunity to come back to NIBB in near future and work under the supervision of Prof. Kiyoshi Naruse and Assistant Prof. Yusuke Takehana to gain as much as knowledge from them.

Dartmouth College, USA
Mikiko Takato
My research project involved studying various signaling proteins by quantitatively manipulating their intracellular abundancies. This was achieved by using a new experimental tool that involves genetically fusing engineered dihydrofolate reductase (DHFR) to a protein of interest, then introducing a small-molecule DHFR inhibitor called trimethoprim (TMP). In the presence of TMP, the fusion protein is stabilized, but in its absence, it is degraded. This allows for the fine-tuning of the intracellular concentrations of proteins of interest. Using this so-called DHFR-TMP protein stabilization system, I studied the phenotypic effect of three signaling proteins, Rac1 (involved in cell motility), MEK1 (involved in cell proliferation), and BAD (involved in apoptosis induction), by genetically fusing them to DHFR and a green fluorescent protein and tracking their intracellular abundancies by fluorescence microscopy.
As a chemistry major and moreover merely an undergraduate, I was completely unknowledgeable and unfamiliar with the field of life sciences when I first arrived at NIBB. However, through this internship program, I was able to learn about and experience biological research first-hand, and even had the opportunity to present my research in a poster session at the annual retreat of the Okazaki Institute of Integrative Bioscience. This research experience also confirmed my interest in attending graduate school to pursue a PhD. I would like to thank Professor Aoki for giving me the opportunity to join his lab through this internship program and also Miura-san, a graduate student in the lab, for her help and guidance in my research project.
My internship experience in the Aoki lab was not only academically enriching, but also (to put it simply) incredibly fun. I felt welcomed in the lab from day one thanks to Professor Aoki’s humorous personality and the warmth and kindness of all of the lab members who helped me all along the way. We would eat lunch together as a lab every day over hilarious conversation, make tacos and spam musubis together for parties with other labs, go out to eat at Cannery Row for the buffet…I am eternally grateful to everyone in the Aoki lab for making my three months here, however brief, so memorable and so invaluable.
Message from Interns 2015

Heidelberg University, Germany
Omnia El Said Ibrahim
One female medaka mutant shows a defect in follicle formation, and some germ cells express the spermatocyte marker Shippo1. in situ hybridization turned up positive for Shippo1 for the mutant line, indicating spermatogenesis is occurring in the mutant ovary. Interestingly, ovaries from one mutant line were also able to fertilize WT ovaries. There is an implication that oocytes might regulate type I germ cell proliferation if the mutants show less oocytes and more type I germ cells. Therefore this was tested through BrdU incorporation of hatchlings and subsequent immunohistochemistry and type I germ cell counting. Unfortunately homozygous mutants were not obtained so this theory could not be tested.
For the other experiment, the purpose was to establish a subcutaneous transplantation system using olvas-eGFP/sox9b-DsRed donor gonadal cells in WT and host and observation of the donor cells after transplantation: how long do the donor cells remain in the host before rejection? As opposed to Zebrafish, using PBS in the water to sustain the transplanted fish does not work well for Medaka. We figured out the optimum conditions to keep the fish alive as long as possible; RO water with streptomycin/ampicillin that is changed every other day and an air bubbler in each tank. Retention of the donor cells depended on the transplantation technique so it was variable, however after the first week most of the GFP signal was gone and the DsRed was retained for longer while also decreasing.
After a lot of direct sequencing and three months later, I can safely say that coming to NIBB and Japan has been one of the most defining experiences of my life. Other than the new techniques I learned in the lab and exciting research I participated in, I was made to feel very welcome by everyone and there wasn’t anything I needed or was lacking. On the weekends I was able to experience what Japan has to offer from beautiful landscapes, mouth-watering foods and fascinating culture. I highly recommend this internship program.

VNU University of Science, Viet nam
Pham Van Cuong
Then I chose laboratory of Assoc. Prof. Kamei, because of his using medaka fish model and heat-shock system, as my undergraduate laboratory. Besides, I wanted to have a chance at IR-LEGO, the technique that using laser to induce expression of heat shock promoter-driving gene in desired cells. Nearly 3 months working in Kamei-lab, I was enthusiastically supported by members of this laboratory and also Bioresource Laboratory. Thank to that help, my experiments (evaluation of heat-shock response and molecular experiments) could turned fluently and be finished with some expected results. I think the obtained results were not significant to me than what I have learned: techniques, experiment manipulation and even working habit. And my questions at beginning also were answered somewhat.
To say something about this program, in unscientific aspect, I feel how lucky I was to join this program: coming to my dreaming country at the first time going abroad, being helped by surrounding people, immersing well in Japanese culture. Thank you for giving me precious experience and memories.

Hacettepe University, Turkey
M. Orkun Çoruh
As a physics engineer, working in a biology laboratory environment was an amazing experience for me. I felt really comfortable in this international environment, with the kind assistance of all students, academicians and support office as well. I had the opportunity to discuss my scientific point of view with everyone, attend to weekly meetings and also an international conference. The atmosphere in the whole institute was very open and supportive, and besides the scientific point of view, there were several social events where I could experience the hospitality and friendship of people while enjoying Japanese culture and cuisine.
The accommodation at the Mishima Lodge was really comfortable with a good stuff. Everything was clean, safe and well organised. The lodge was really close to Institute, and also to the city center.
My time in this research internship was really efficient in scientific and also social point of view. Besides learning new methods and getting acquainted with the point of view of world class biology scientists, I had the opportunity to enjoy the well protected nature. Ultimately the time I spend in Okazaki was rewarding and valuable for me.

Universidad Nacional del Santa, Nuevo Chimbote – Peru
José Alexander Carranza Luna
I was an intern in the Laboratory of Molecular Genetics for Reproduction, which is led by PhD. Tanaka, Minoru from 4th December 2015 to 1st March 2016. I came to the NIBB after obtain a scholarship from the Peruvian government, specifically for the National and International mobilization program in science, technology and innovation from the Consejo Nacional de Ciencia, Tecnología e Innovación FONDECYT-CONCYTEC (Peru), in order to do a: “Training in techniques for sex determination and differentiation in fish” and contribute to the project: Sex Determination and differentiation in Arapaima gigas “Paiche” (the second freshwater fish largest in the world), which is executing in the Laboratory of Genetics, Physiology and Reproduction in Peru, which I take part.
During my stay at the NIBB I could learn various techniques such as: in situ hybridization, immunohistochemistry, confocal microscopy, genome editing by CRISPRcas9, probes designing, bioinformatics software and others, by using the Japanese fish Oryzias latipes “medaka”. I studied about the germ cells biology, how the gonad is form and what genes are important by analyzing several mutants strain. Since the germ cells are a set of very important cells because they are responsible to transmit the genetic information to the next generation and also have the ability to form a whole individual, I am very interesting for continue doing research in this fascinating field.
I would like to thank PhD. Tanaka, who gave me the opportunity to join the lab and for his suggestion in the experimental process in the Paiche project in Peru. Likewise, I would like to thank to the lab members: Shiraishi, Sakae, Fujimori, Kikuchi, Otake, Watakabe, Suzuki and Kinoshita for helped me with the experiments, for their kindness and their friendship. Specially, I would like to thanks to Nishimura-san, who was my mentor along this 3 months, thanks for his time, dedication, for the knowledge that shared with me every time, for introduce me to the Japanese culture and for his sincere friendship. Also, I would like to thank to the members of Miyanari's Lab who were friendly and kind.
Along this 3 months, I could visit some touristic places as Okazaki Castle, Takeshima Shrine, etc., which helped me to know and understand a more details the Japanese culture. Also, I could taste several typical food (Yakiniku, was one of the best) and enjoy incredible moments in this beautiful and fascinating country.
Eternally grateful.

Mahidol University, Thailand
Rawinsak Phumthananiwet
I joined Dr. Kawade Kensuke’s laboratory which focused on plant development and physiology. This laboratory has been studying metabolic systems that coordinate with developmental progression in plants, especially in Arabidopsis. We are interested in ANGUSTIFOLIA3 (AN3) which is a signaling molecule involved in the control of proliferation of epidermis and mesophyll in leaves.
I had an opportunity to do some experiments including root growth kinetic comparison between AN3 wild type and an3 mutant, expression pattern of AN3 encoded GFP and AN3 promoter-ß-glucuronidase (GUS) reporter line in order to prove and illustrate expression of AN3 in root. The results from these experiments were consistent with the previous studies. I also had a chance to do root confocal analysis, leaf area and cell size experiment.
The Arabidopsis plants used in these experiments were grown in our laboratory. Root length and expression of AN3-GUS and AN3-GFP was observed by using confocal microscopy. The photos were captured in order to measure cell size by using Image J program. The area of Arabidopsis leaves were also measured by using Image J program and leaf palisade mesophyll cell number was calculated from cell size and leaf area.
This internship program is very good for students who are interested in Biology Science and would like to enjoy Japanese culture including working style, life style and so on. There are many new experiences you can receive here. Especially, for students who would like to be a master or Ph.D. student in Japan, This internship would be a great choice.

Okayama University Office in Viet Nam, Vietnam
Nguyen Thi Hong Dung
I had great internship. In one month, I studied a lot of skills and knowledge. Surely, this program was a rare chance for anyone who wishes to experience the professional study and research environment at NIBB. Through the internship program, with the guidance from professors and lab mates, I practiced research skills, including: Techniques, experimental operation, data recording and analysis, methods of data reporting, etc. Besides specialized knowledge and skills in scientific research, I learned the work ethic of scientists in one of the fore-running scientific labs of Japan – one of the leading countries in terms of discipline, enthusiasm, and hard work. During my internship, my lab mates not only helped me in my research but also in learning Japanese. They were very friendly and kind. I would like to thank all laboratory members: Naomi, Watakabe, Sakae, Furimoji, Nishimura, Kinoshita, and Otake. I would like to thanks Ms.Mariko Kikuchi, she was very thoughtful and kind, she helped me a lot in research work as well as daily living. With her helping, I was always very happy when I stayed in Okazaki city. Especially, Dr.Tanaka was very kind with me, he gave me a feeling of comfort. I will keep these great memories in my heart.
These experience would be very valuable to me, will help me adapt to and successfully complete my one year of master’s program at Okayama University as well as leading me to become good researcher in the future. I would like to thank Dr. Tanaka Minoru and NIBB for granting me this opportunity
Message from Interns 2014

University of Pécs, Hungary
Pálfalvi Gergő
In this time the reason of my visit was to investigate some in silico data from C. follicularis’s genome which is connected to the nutrient acquisition and to the carnivore syndrome, too. For this, first I learned the basic use of UNIX and shell scripts on NIBB’s super computer. After that I learned how to make a phylogenetic tree for homologous genes starting with BLAST search and finishing with the gene tree. I examined the plant’s messenger RNA (mRNA) transcriptome data and found differentially expressed nutrient transporters which can be good candidates for genes involved in carnivore syndrome. Now I can use advanced bioinformatic tools and I can start my own project at Hungary with microRNA (miRNA) transcriptomics which is highly connected to the project of Professor Hasebe and Mr. Fukushima.
There was a second aspect to my visit this time. In December I will take the examination for SOKENDAI and I would like to investigate the pitcher development in the future. To start this project I optimized in situ hybridization protocol for C. follicularis including tissue fixation, embedding, serial section making and in situ hybridization. This time I have succeeded with Histone H4 gene which is a good indicator for actively dividing cells. Now I could describe some part of the developing pitcher’s division areas but my data is not enough for conclusions yet. To describe the development phases of the pitcher leaf, I made serial sections from shoot apex grown on pitcher inducing- and also on flat leaf inducing environments and stained them with Toluidine blue. Now I am processing the data, but I can say it looks like the determination of leaf type occurs in a very early developmental stage of leaf primordia – which is opposite with previous thoughts and the development is quiet different from other non-related carnivorous plant’s pitcher leaves, such as Sarracenia purpurea (Mr. Fukushima’s project was the development of the pitcher leaf of S. purpurea).
On the extracurricular programs, I am wondering about the Japanese old and rich culture and it was a great opportunity to get to know with it. The atmosphere in the laboratory was always great and I have had opportunity to discuss with and get inspired by magnificent researchers.
After this trip I can say I am richer with a lot of knowledge, inspiration and great friends, in addition to a lot of experience about international collaborations.
I would like to thank you this opportunity and in the future I would like to join Professor Hasebe’s team to examine the evolutionary development of the pitcher leaf of C. follicularis.

Shanghai Jiao Tong University School of medicine, China
Dilukshi Chinthani Perera
Germ cells are a type of biological cells that involves in reproduction. In many animals; the germ cells originate in the primitive streak (structure that forms in the blastula during the embryonic development) and migrate via the gut of an embryo to the developing gonads. There, they undergo cell division of two types, mitosis and meiosis, followed by cellular differentiation into mature gametes, either eggs or sperm.
Germ cells play an essential role in Sex differentiation in Medaka. It is required for ovarian formation and also Germ cell deficient gonads develop a testis-like structure.
In vertebrates there are two modes for the determination of sex
1.Genotypic Sex Determination (GSD)
2.Environmental Sex determination (ESD)
•Mammals and Birds follow GSD.
In medaka sex-determining gene dmY/dmrt1y has been identified. My studies were mainly focused on GSD, two important genes.
At the beginning of the experiment I had to dissect Medaka lava to extract gonads which was a very interesting activity. I enjoyed collecting fish eggs at Myodaiji. Performing micro surgery was where I used my skills as a medical student who had assisted surgery at hospital. It was an opportunity to exchange knowledge with the PhD students at the lab. Therefore, my overall experience was invaluable. I certainly had a very useful time with Professor Tanaka and his team.

Bangalore University, India
Shashank Kumar
I successfully completed the project and learned many advance techniques, which are going to be very helpful for my future research work. I am sincerely thankful to Mii San who has taught me different techniques, which are used in recent molecular and developmental biological research.
My primary objective was to experience basic DNA construction, which is obtained by double digestion of two restriction enzymes. The DNA were isolated and purified by gel purification method. We have used two specific type of strain (mutated and non-mutated). After the digestion we have done the gel purification and isolated the specific gene. This specific gene was then transformed to the competent cells of E.coli strain. After the incubation period the transformed E.Coli cells were observed and colonies were picked for mini culture, for the specific transformation we have done the colony PCR that indicate the successful transformation of the cell. All samples were successfully transformed and the result was obtained successfully. Apart from this Mii San has taught me about the Confocal Microscopy and Microinjection of XWnt mRNA in Xenopus laevis.
As a Biotechnology background, these experiments were totally new for me but I sincerely thanks to Mii San for guiding me in each and every step. He explained me all the basic principle for all the experiments and taught me the minor specification in the experiments, which is very useful for my future research. Even I am sincerely thankful to Sir Prof. Shinji Takada, Mii San and all the lab members, which has helped me a lot for my presentation. During my stay, Sir Prof. Shinji Takada and all lab members has given me a very warm welcome dinner party at an Indian Restaurant for that I am really thankful to them. As it is known to the world Japan is very rich in culture, I had a dream before to visit Japan and the dream came true due to this internship program. Even my all lab members were very keen to know about the Indian culture and I have told them about the Indian culture.
At last I would like to thank to Ritsue Takahashi sir, for guiding me from India to Japan. I would also like to sincere thanks to Ukai San for the administrative work and helping me a lot during the stay in Japan. I would like to sincere thanks to NIBB and Sir Prof. Shinji Takada for granting me this opportunity. I would also like to thank my all lab members for caring me a lot. I am also delighted for spending my whole tenure at Mishima Lodge; the facilities were very comfortable for making my stay delightful.
This Internship was very important for me to become a good researcher and a good human being for my future. I sincerely admire this country with most honest, hardworking and helpful people around there. I really want to continue my further research work and I hope I will get the chance again to carry out my research in this lab. Thank you very much.
Message from Interns 2013
Huazhong Agricultural University, China
Nan GU
During this period, I mainly focused on studying the function of Topoisomerase 1 (Top1) in the moss Physcomitrella patens. TOP1 has important functions in DNA replication, transcription and chromosomal segregation. As we know, Physcomitrella patens is an emerging bryophyte model organism for various aspects of plant evo-devo resesrch and Prof. Hasebe’s laboratory has established a well accepted and simple research system to study the phenomenon of stem cell formation from wounded differentiated leaf cells of P. patens, so I decided to study the function of TOP1 using this system to further our understanding of the Topoisomerase 1, and gain an evo-devo view of this enzyme. By this chance, I had an intuitive understanding of P. patens and its advantage in my research. I also learned many useful and advanced techniques, like construction of a targeting vector, generation of mutants, culture and storage of protonemata and gametophores, observation of protonemal development by a time-lapse system and so on. At last, I smoothly finished the work as planned and I’m delighted to get some unexpected experiment data.
I feel so lucky to be an intern of the NIBB. Here, the relaxed and open scientific research atmosphere makes me feel comfortable and I am impressed by the efficient experimental system and rigorous academic attitude in NIBB. Happily, Hasebe sensei offers me another chance to stay in his lab for a longer time as a special collaboration research student from January 2014 to March 2014.
Apart from research work here, during week-end holidays I had a visit to Tokyo in October and Kyoto in November and some scenic spots around Okazaki. The beautiful scenery and hospitable Japanese people were imprinted on my mind. In December we had an interactive communication visit to Ryukai junior high school and we enjoyed a memorable afternoon with those outgoing, lively and cheerful students. I feel much honored to experience the local conditions and customs of Japanese.
Finally, I would like to thank NIBB and Prof. Hasebe for giving me this opportunity. This experience in Okazaki strengthened my conviction in scientific research. I’d also like to thank to all members in Hasebe lab for their help in my work and life during my stay. If it is possible, I want to do some further researches in Hasebe sensei’s lab.

University of Dhaka, Bangladesh
Manirul Haque
I have successfully completed the experiments and also learned many advanced techniques. To detect laminin I have to learn Immunohistochemistry which was first time for me. I have done this experiment using 18 dph (day post larvae) Olvas transgenic medaka. GFP ( Green Flurocent Protein) was used to detect the gonad which express green color in Germ cell under microscope. There is a specific antibody (anti laminin antibody derived from rabbit immunology) to detect the laminin signal in germinal cradle. Laminin signal indicates the basement membrane which generally stays on the dorsal side of the ovarian cavity and also it remains around the outer side of the follicle and specifically divides the inner and outer shell of the follicle. I had done this experiment three times and at last I was able to detect strong laminin signal. I have also done another experiment that was detection of laminin in matured gonad of hotei medka. Interestingly, We found oocyte like structure in XY male hotei gonad along with spermatocyte. Besides, I learned XY typing (Genotyping) of hotei (germcell excess medaka ) , DNA sequencing, microinjection , using of confocal microscope and so on.
Working on such types of experiments, I had just found that this field was more convenient than learning just a theory. I specially thank Dr. Chika Fujimori . She was really very amiable and helped me in every aspect and explained me every problem very well. All the other lab mates ; Toshi , Yuta, Otake, Kikuchi were really enthusiastic to talk and make fun with me. Every lab member was very open and ready to help. I was so overwhelmed by such nice lab environment. I am also very grateful to Manami and Sujuki for their nice cooperation with me.
During my one month stay I visited Nagoya port Aquarium, Okazaki castle and other temples. I enjoyed the Japanese culture a lot. All the lab mates were very keen to know about Bengali culture which fascinated and touched me a lot.
At the last I would like to thank NIBB and Dr. Tanaka for granting me this opportunity. Also I would like to thank all the lab members and staffs for caring me. I am also delighted spending my whole month in Myodaiji lodge. The facilities were fantastic for making my stay comfortable and enjoyable. I would also like to share some photographs which are life time memories for me which I have attached to the file.
This internship was very important for me to become an efficient researcher in future. I enjoyed this beautiful country with most honest, hardworking and helpful people around there I ever saw. I really want to continue my further research in this area and hope I will get a chance again to work at NIBB particularly in this lab. Thank you very much !

University of Pune, India
Bhagyashree Nandkishor Swarge
I have heard experiences of my friends who had been selected for the same program and worked in NIBB previously. They told me about the state-of-the art technology at NIBB, the well equipped labs, the research, Japanese culture, people and so I was very excited to come here.
Right from my selection to arrival in Japan I was guided very promptly and sufficiently regarding every query by Ms. Ritsue Takahashi. Everyone in my lab was very friendly and helping. My lab mates not only helped me in my research work but also in planning my weekend trips to other famous places in Japan and arranging train tickets. I am very thankful to all of them.
As I am from microbiology background this internship work and the techniques used were completely new for me. From this internship work, I have learnt to grow algal cell in controlled environment. I have worked on the transformation of unicellular eukaryote by electroporation method. My research focused on the nuclear transformation of Chlamydomonas and to check the transformants for exogenous expression. Dr. Ryutaro Tokutsu guided me and clarified my every doubt patiently. I am also thankful to Konomi san to teach me electroporation technique.
During this period I also got opportunity to attend Journal Club, where the recent published work in the field of photobiology was discussed, which helped me to understand and broaden my ideas in this field. I would like to thank NIBB and Dr. Jun Minagawa for granting me this opportunity

Capital Normal University, Beijing, P.R. China
Liu Meng
Dec.11 A.M: Dr. Takeda introduced the labs to me and took me a tour around NIBB.
It is really an amazing lab. This is my first time to see so many great facilities in one lab. Also Dr. Takeda showed me some confocal systems and culture systems in other labs. The big green house impressed me so much.



Dec.11 P.M: Dr. Takeda showed me how to observe AM fungi’s hyphae using a fluorescent microscope and how to use a confocal system.
We picked some transgenic seedlings and stained by DAPY. Then we observed them under fluorescent microscope. And Dr. Takeda showed me that we can use confocal system to detect the Ca spiking. (I’m sorry but I left the pictures on the microscope’s computer.)
Dec.12 A.M:The seminar of Prof. Kawaguchi’s lab.
Dr. Sayano gave us a great presentation about < SCFKMD controls cytokinin signaling by regulating the degradation of type-B response regulators>.
I knew a little about SCF complex as well as the cytokinin signaling pathway. But I didn’t know their connection until listen to Dr. Sayano’s introduction to this paper. Cytokinin receptor binds to the CHASE area of the activated receptor and let it trans-autophosphorylate. Then the activated receptor phosphorylates the subfamily AHP and make it move from the cytoplasm to the nucleus. The interaction of the active AHP and ARR in nucleus will transfer the phosphate group to the accepted domain of ARR and the type-B ARR release from the nucleus. The dephosphorylated AHP can return to the cytoplasm and be phosphorylated again. Type-B ARR can combine to multiple cis-acting elements of the target gene’s promoter then activate the expression of type-A ARR, which will interact with other regulators and change the cell. Type-A ARR activation provides a negative feedback regulation of its own expression. The author found out that KMD genes encode F-box protein. Their mutants are sensitive to the cytokinin while overexpression the KMD reduces the sensitivity. And the change of ARR at expression level consistent with this. They also proved that KMD have direct interaction with type-B ARR. Based on above they proposed a model of SCFKMD-mediated regulation of cytokinin signaling pathway that SCFKMD target the type-B ARRs for degradation through the ubiquitin proteasome pathway to control the quantitative of available type-B ARRs.
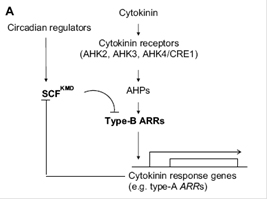
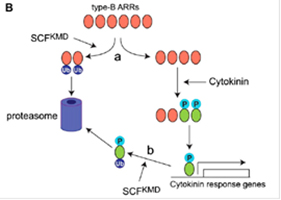
Dec.12 P.M: Dr. Suzaki showed me some mutants of Lotus japonicus in our lab. Then he taught me about the LacZ staining and Dr. Tanaka taught me how to mate the flowers between different ecotypes.
I read some papers before and had a little confusion that how did they find the difference between the WT and mutants, the nodules are so small in a plant? But when I saw them under a microscope it had a big difference. Different mutants have their special characters, such as the numbers of the nodule, the color and the size. Even some mutants have the same number of nodules compared with WT but they are localized in a wider zone in root. I got a visual impression of several mutants including har1 daphne etc.
What I think the most interesting thing is the nodule’s color. I didn’t notice that they’re red when I read the papers. Dr. Suzaki told me that this is caused by Leghemoglobin (plant hemoglobin). I checked some reviews that it is first found by Kubo in 1931 and he proposed that Leghemoglobin may play a role in oxygen transportation and plant assimilation. It can keep the hypoxic environment surrounding the nodule that it only needs 7-11nmol/L O2 in the cell. So the Leghemoglobin will protect the oxygen-sensitive nitrogenase and also promote the spread of free oxygen. Ott proved this in 2005. He silenced the sHb gene in Lotus japonicus and found that free oxygen increased in nodule and nitrogenase been damaged causing the loss of the nitrogen fixation.
Under Dr. Tanaka’s instruction I tried some pollinating experiments. She told me that Lotus japonicus has two main ecotypes that they usually use: Miyakojima and Gifu. Sometimes they need to graft them to purify the genetic background. It is very hard to do mating on such small flowers. But finally I began to know what kind of flowers is more appropriate for pollinating. After killing so many flowers Ms. Tanaka told me that this one probably can survive.


Dec.13 A.M: Dr. Suzaki and I checked our staining result and observed the infection threads using confocal system.
Our staining is good. Under the microscope I began to understand the difference of the infection thread between WT and other mutants. Some mutants don’t have threads at all but some have threads still cannot form the nodules. Unfortunately I left the pictures of them on the confocal computer.
The fortunate thing is that I learned two tips about the staining. When I do the GUS and LacZ staining I always vacuum excessively because I used to keep the machine running. Now I know that I can stop the vacuum but keep the container sealed for a while to hold the pressure. It will take the time a little longer but won’t hurt the sample especially some vulnerable tissues like the leaf discs. And according to our manual book that the phosphate buffer should be prepared freshly and I have to mix the Na2HPO4 and NaH2PO4 as 3.9:6.1 every time so it wastes time and reagents. During the internship I found them just use the premixed buffer and has no effect on the results. I used these experience after I got back and works well.
Dec.13 P.M: Some students introduced their current research to me. Dr. Tanaka instructed me about grafting.
This is my first time that using a microscope to complete a grafting experiment. It is really a delicate work. But I think I’ve found some tips about that in the end after some practice. The cut to the stock doesn’t need very deep and the scion should better have a sharp angle. No water should be left at the junction part. So it is better to keep the stock a little higher than the edge of the paper. And some of my work survived after I left Japan.



SUMMARY
The whole Internship trip is so fantastic. It is full of “first time” to me, the first time coming to Japan, the first time using kinds of microscopes for study (usually I just use them once or twice in a year due to my current major and study) and the first time I accomplishing mating and grafting on such small plants etc. Three days experiments gave me some direct impressions of Prof. Kawaguchi and his team’s work. There are two regretful things to me. One is I didn’t read enough latest papers before I go to Japan so I didn’t catch up some of their introduction of their current study. Another is even though I can speak and understand English but I’m not accustomed to write the notes or records in English while doing experiments. It is a habit I need to prove.
Overall I’d appreciate NIBB and Prof. Kawaguchi offering me this chance. Thank you for your thoughtful arrangements. Thanks for Mr. Takeda, Mr. Suzaki and Ms. Tanaka introducing and instructing a lot of experiments for me. Thanks for Yoro-san, Fukuhara-san, Nishida-san and Sasaki-san accompanying with me all the time. And thanks for all the people in Prof. Kawaguchi’s team that gave me so much help during my stay in Okazaki.
Message from Interns 2012

Swami Ramanand Teerth Marathwada University, India
Ravi Prabhakar More
Really it would be my pleasure to have the chance to be an international intern of this respectable institute in National Institute for Basic Biology, Okazaki city in Japan. My work in the laboratory was very interesting and enthusiastic in which I was able to combine metagenome with informatics in addition to implement scientific programs such as “Study of Horizontal Gene Transfer events in microbial community: Metagenome sequence data as a model” by using highly qualified software RECOG and NIBB high performance computing facility.
My internship helped me to gain a lot of experiences and skills in Bioinformatics fields especially in high performance computing and comparative genome/metagenome field as well as being able to communicate with different culture and society. During the internship period I also delivered seminar on “Mining the Metagenome from CETP: Prediction of microbial community and novel metabolic network”. This seminar gave me nice opportunity to interact and discuss scientific idea with NIBB scientists.
Apart from my research activity I thoroughly enjoyed my stay at NIBB, Japan especially the visits to Mount Fuji and Kyoto temples. At the last I would like to thank all Genome Informatics laboratory members to make my stay so comfortable and joyful whereas I would personally like to thank Prof. Dr. Ikuo Uchiyama for proving the nice ideas to work upon. I am also highly indebted to Dr. Hirokazu Chiba who seemed to have solutions to all my problems. I would also like to share some photographs which are life time memories for me which I have attached to the file.

The Modern College, India
Sneha Mahalunkar
My work majorly focused on the DNA methylation patterns of the differentially expressed homologous gene sets derived by tetraploidization during evolution of Xenopus to examine whether DNA methylation contributes to the multifunctionality of duplicated genes in various stages of Xenopus laevis embryos.
According to the cluster analysis of differentially expressed genes by RNA sequence of embryos from several developmental stages, hundreds of homologue gene pairs were previously identified as gene sets that show two genes displaying temporal expression profiles in early (before MBT) and late (after MBT) stages. Among them, the pair of X1_pdlim7.a and gene X1_pdlim7.b were chosen for the analysis of epigenetic status, namely the methylation of DNA because they show a reciprocal temporal expression pattern during development (Data Provided); 7a is expressed at high levels in the early stages and low levels in the late stages and on the contrary, 7b is expressed at low levels in the early stages and high levels in the late stages.
Since the gene regulatory sequences of X1_pdlim7.a and gene X1_pdlim7.b were thought to be near the 5’ UTR on the Xenopus genome, the 5’ most end region of the 5,UTR regions were chosen for the DNA methylation analysis by bisulfate sequencing.
Briefly, genomic DNA obtained from Xenopus embryo was treated with bisulphate conversion reaction followed by a PCR amplification of the 5’ UTR region and sequencing. The sequencing results show whether CG nucleotide sequence within the 5’UTR regions is randomly methlylated.
I first amplified the Xenopus genomic DNA previously prepared by Dr. Miho to bisulphate conversion of DNA. I amplified the 5’ UTR sequence of X1_pdlim7.a and gene X1_pdlim7.b by using PCR. These amplified sequence were analyzed by using agarose gel electrophoresis. Out of the 8 samples (that is amplified for two primer sets; X1_pdlim7.a and gene X1_pdlim7.b and for 4 templates; stage 9 and stage 12.5,raised by two independent individuals for each ones amplified with a 4-primer set b gave expected result of 230bp of intensed band still we used the 8 samples for further analysis.
All the 8 PCR products were ligated into a vector and transformed into E.coli cells by using the Strataclone PCR cloning Kit. The transformed colonies were selected by blue white screening method. We selected 8 colonies from each plate (Plate 1,2,3,4,5,6,7 and 8) and conducted colony PCR of the selected colonies. After colony PCR 10 microliter aliquots were electrophoresed to determine the length of the inserts.
Inserts roughly estimated to be the desired insert 230bp were selected and subjected to ExoSAP method to remove the unused primers. The inserts were sequenced by DNA sequencer and resulting sequence data are analyzed by BiQ Analyzer software to identify methylation site.
The result we got was that the 5’ UTR of the genomic DNA derived from stage 9 amplified by a primer set ‘b’ was methylated (analyzed by BiQ Analyzer), which was an expected result as the gene X1_pdlim7.b is a late expression gene and therefore to suppress the gene expression before the onset it should be methylated in its early stages including stage 9.
After the completion of my work allotted I was filled with thorough knowledge on how epigenetics work can actually be carried out. I am also very much motivated to conduct such kind of work with more thorough understanding in this field. The best thing encountered during my work was to learn the application part of Nested and Colony PCR which I knew only theoretically. I am also glad I got to learn Sequencing and its data analysis which was also a new experience for me.
I had a wonderful experience during my course period and I am deeply thankful to Professor Naoto Ueno for giving me such a great opportunity. I am also very much thankful to Dr. Miho san who guided me throughout my work and was so very kind to me for explaining me the entire work that had to be done. I am also thankful to Dr. Suzuki who explained me his work on calcium influx regulated neural tube closure development. I am grateful to all the PhD students namely Ms. Miyagi san, Mr. Hara san and Mr. Hayashi san who could take out some time to explain me their work with so much of patience. Last but not the least I am thankful to all the staff members especially Ms. Toyoko san who took very good care of me and made me feel comfortable. I am thankful to NIBB for giving me the opportunity to understand the research culture as well as to experience the Japanese hospitality in Okazaki.

University of Pune, India
Tushar Suhas Khare
I have successfully completed the work assigned to me and have got well experienced with advanced techniques. I learned how to grow the Chlamydomonas cells in proper culture system by maintaining the exact conditions. I got opportunity to handle some new instruments such as Light meter and Chlamy-spec. I performed the experiments which include SDS Gradient PAGE, Immunoblotting, Pulse Amplitude Modulation Spectroscopy for Non-Photochemical Quenching measurements. During internship, I had found this field more interesting than learning just a theory. Dr. Ryutaro Tokutsu and Dr. Kenji Takizawa helped me in every aspect and explained me every problem very well. All the other lab mates and assistants in lab were really enthusiastic to talk with me. Every lab member was very open and ready to help. I was so impressed by such healthy lab environment.
During my one month stay I visited Nagoya, Kyoto and Tokyo. I also roamed around in Okazaki. I enjoyed the Japanese culture a lot. I also celebrated most important Indian festival DIWALI with my lab mates. All the lab mates enjoyed that and were very keen to know about Indian culture which fascinated and touched me a lot.
At the last I would like to thank NIBB and Prof. Jun Minagawa for granting me this opportunity. Also I would like to thank all the lab members in Prof. Minagawa’s laboratory for everything. I am grateful to Mishima lodge facility for making my stay comfortable and joyful. I would also like to share some photographs which are life time memories for me which I have attached to the file.
I wish all the very best for every lab member for their future work. I am inspired by the people I worked with. I hope to have another opportunity to come back NIBB at future. If I get such chance, I will be glad to acquire it. Thank you !


Karlsruhe Institute of Technology (KIT), Germany
Alexander Weichsel
Wnt signaling has important roles during development and in many diseases. Many studies have identified several factors that are required for the secretion of Wnt proteins, however the secretion pathway and the establishment of the extracellular distribution are yet unknown.
I have got the possibility to perform a project about Wnt secretion process. The objective of this project was to visualize the secretion process of Wnt ligands by using different cell culture techniques. My supervisor for this internship was Mii-san. He introduced his work and we discussed my detailed schedule. Everything was well planned and he taught me many techniques. I learned how to dissect mouse to remove brain and collect hippocampal tissue from mouse embryos. This was exciting for me, because it was the first time I worked with mouse. Before, I only gained experience with Xenopus and Zebrafish and during this period I learned a lot about mouse development, which was very useful for me. Furthermore I did cell transfections, several immunostaining techniques, live cell imaging and learned new techniques which are important in the field of molecular biology. Finally I was able to establish a technique for live cell immunostaining, which can be used in future experiments.
My accommodation at the Myodaiji Lodge was close to the Yamate campus and easy accessible. I did not had any trouble during my stay and I am very grateful for that.
Finally I thank NIBB for giving me this great opportunity to develop my scientific skills and experience Japanese hospitality. I am deeply grateful for this opportunity and want to thank especially Takada-san and all lab members for their hospitality, scientific guidance and the great atmosphere. Thank you very much!

Capital Normal University, China
Ji Zhongzhong
Leaf cells of the moss Physcomitrella patens can be reprogrammed to protonema apical cells with stem cell characteristics by leaf excision. Previous works in Prof. Hasebe’s laboratory on the reprogramming have identified a transcription factor STEMIN, of which overexpression in an intact gametophore can induce reprogramming of gametophore leaf cells into protonema apical cells without excision. However, how STEMIN induces reprograming remains still elusive. During my internship program, to address this question, I investigated spatiotemporal expression patterns of STEMIN in excised leaf cells using the GUS knock-in line and changes in expression of genes involved in reprogramming after the STEMIN induction with advanced techniques.
I successfully completed the works described above and obtained some experimental data. At the last day of my internship program, I gave a short talk about these results and discussed with Prof. Hasebe and his lab members. It helped me to understand plant stem cell formation.
During my staying in NIBB, everything went well; everyone here was friendly, enthusiastic and accommodating, I felt very lucky for joining the Prof. Hasebe’s lab. The NIBB offered many seminars for students and stuffs, it is good platform and helpful for enhancing communication and open mind to researchers here. Apart from research works here, I had opportunities to visit to Fuji Mountain and Inuyama castle with lab mates, it was great experience to me. Finally, I want to thank you Prof. Hasebe, Dr. Ishikawa, and all of lab mates in Prof. Hasebe’s lab, thank you for your help during my stay in NIBB.


Central Institute of Fisheries Education, India
Aritra Bera
Sufficient oxygen supply is a key element for reproductive success in animals as well as in fishes. Hypoxia (generally dissolve oxygen < 2.8 mg/l) in natural water is a worldwide problem now-a-days. Low oxygen content due to aquatic pollution and global warming in marine water and freshwater is affecting reproduction and endocrine function to a broad range of fishes every year. But effect of hypoxia on germ cells is not well understood in fish model till now. So we exposed adult male and female medaka (wild type, Sox9b-DsRed/Olvas-EGFP & Aromatase-EGFP) to chronic (24h daily) and cyclic (12h daily) hypoxia (1 mg/l dissolve oxygen) for 24 days to understand hypoxic responses of gonadal germ cells and somatic cells by quantifying VASA, Sox9b, Aromatse, BrdU and Caspase-3 signals through Immunohistochemistry in confocal microscopy. We also studied brain aromatase and caspase-3 activity under hypoxia with relation to BrdU positive neural cell proliferation under hypoxia and normoxia. Hypoxia (chronic and cyclic) was enhancing apoptosis and inducing mitosis (indirectly) in germ cells and somatic cells by expressing Caspase-3 and BrdU signals respectively in both sexes. Hypoxic gonads resulted in low GSI were expressing more VASA cells. Spermatogenesis was disrupted in hypoxic male may be due to less expression of aromatase in testis. We also had performed whole mount In situ hybridization of HIF-1 alpha in ovary showed HIF-1 alpha expression in immature oocytes under hypoxia (cyclic and chronic). Male brain showing caspase-3 positive cells and negative aromatase expression was more prone to hypoxic damage than female where as in female hypoxic brain showed more BrdU signal. Similarly we exposed 5dpf Sox9b-DsRed/Olvas-EGFP & Aromatase-EGFP embryos to chronic hypoxia (0.5 mg/l) upto 5dph (7 days exposure). Hypoxia was showing enhanced germ cell apoptosis in male and female 5dph juvenile than their normoxic counterpart. Male juvenile expressed more HIF-1 alpha in gonad than female whereas female juvenile were showing enhanced gsdf expression during q-PCR analysis.
This internship was very important for me to become an efficient researcher in future. I liked the sense community and the friendly atmosphere in NIBB: not only students but also the staff and professors were outstandingly supportive. Furthermore, a nice lodge program and well organized student organization made me a very smooth transition to the life in Okazaki during my internship. Each and every day was very precious for me and I have learned many new things during this time. I am really thankful to Dr. Tanaka for his unforgettable help, valuable guidance and moral support throughout the internship period towards me. I really want to mention the help I got from my lab-mates specially Yu-ta-Sakae and Toshiya Nishimura during my experiments, unless them it will not be possible to execute it smoothly. I had enjoying this beautiful country with most helpful people around there I ever saw. I really want to continue my further research in this area and hope I will have a chance again to work in NIBB in future.
University of Pune, India
Krunal H Patel
During my internship tenure I got an opportunity to attend the journal club presentation of research papers in the field of Neurobiology. This helped me boost up my understanding in variety of concepts in Neurobiology and gave me an overview of current interests of various groups in Neurobiology research at NIBB. I would like to thank my mentor Assistant professor Nakayama who guided me politely and patiently during my working tenure out of his busy schedule in the laboratory. I would also like to thank Matsuda san for her enduring support during my stay at the laboratory.
I had an opportunity to see Okazaki and its cultural heritage, which amazed me with its beauty and greatness. Every day I learned some new technique and last but not the least, I had experienced a very delightful scientific environment with right proportions to prosper as a doctoral student in future. I wish my best regards for future success to members at NIBB.
Message from Interns 2011

Justus Liebig University Giessen, Germany
Yousef Yari Kamrani
I liked the sense community and the friendly atmosphere in NIBB: not only students but also the staff and professors were outstandingly supportive. Furthermore, a great lodge program and well organized student organization made me a very smooth transition to the life in Okazaki during my internship.
I am inspired by the people I worked with and I hope to have another opportunity to come back NIBB at future.

Australian National University
Pichestapong Phannipha

Tamil Nadu Agricultural University
Veera Ranjani Rajagopalan
I would like to thanks NIBB for giving me this golden opportunity. Each and every day was very precious for me and I have learned many new things during this time. I am much honor for developing my professional skills of the scientific research which is needed for every basic science student. I was very impressed with the multidisciplinary research areas in the neuronal cell biology lab. Hence I feel to study and complete my doctorate career in this innovative area.

National Chemical Laboratory
Kulkarni(Vaman), Tejaswini
Right from my selection, upto my joining for the program; I had been informed promptly regarding my every issue/query from sensei as well as lab mates which made me comfortable. My work schedules and my stay were well planned. Everybody in lab was really enthusiastic to talk with me. I was so impressed by well-equipped lab and the most important was healthy lab culture.
My mentor for this internship was Chensan. She is doing her post doctorate. Although she was piled up with her own work, she had taught me each and every technique with perfection by taking time from her busy schedule. As I am originally form Microbiology background, internship work and techniques were new for me. But I had learned this subject as my theory paper during master’s course. During internship, I had found this subject more interesting than learning just a theory. I had done experiments regarding expression pattern of Wnt 3A in zebrafish embryos and overexpression of Wnt 3A mutants. I had got good practical knowledge of important techniques of molecular biology like DNA and plasmid purification, in situ hybridization, probe preparation, microinjection etc. which are key techniques in molecular biology field.
Apart from work I had enjoyed my whole stay with my lab-mates. I had got good well-come party as well as fare well party by my lab. Everyone in lab is really friendly and helping nature. I have got good friends from this lab. Internship program at NIBB was golden opportunity to get acquainted with respect to knowledge as well as to know more about Japanese work culture. If I get a chance to come to NIBB for further studies, I will be glad to acquire it.

American University in Cairo
Yehya Mohamed Shawky Ismaeil
Actually I was so insist to come to NIBB I applied before for the internship and finally they accept me, when I received the acceptance email of the internship it was like a dream for me , wow I will go to japan especially NIBB at okazki .
My project is with prof NONAKA its about “ nodal cilia flow ( detection by high speed camera )“ . really this is very interesting branch of science and also it is very useful for me and my experimental work so much.
Really , during this three weeks I learned a lot of new techniques and work at great scientific facilities , and very cooperative, respectful and polite peoples “ simply they are Japanese “.
One of my next dreams is to join the Phd course here , I will do my best to accept again for this great institute and to visit this most beautiful country I have ever seen before “ Japan “.
Thank you so much for all the people who help me to come here and help me to success in this internship.
Thanks ALLAH
Thank you my mother , father and my brothers
Thank you prof. Nonaka
Thanks Egypt.

Academia Sinica
Chen-Wi Hsu
During the one-month staying in Prof. Tanaka’s laboratory, I focused on learning techniques used in studying gonad development in medaka. First, to realize the gain-of-function effect of genes of interesting during early gonad development, we must generate the transgenic fish that certain genes can be induced at specific time point and in specific tissue. The heatshock system developed by Tanaka’s laboratory is a powerful tool for gene induction. Therefore, I learned the skills about whole fish or partial tissue heatshock in adult medaka or larvae. The whole fish induction was performed by water bath. The results showed that the GFP fluorescent can be induced both in adult and larvae. The partial tissue induction was performed by probe or IR-LEGO. Although I still need more practice, GFP in some of tissues in adult medaka and larvae were induced successfully. To realize gene expression differences between male and female more effectively, gonadal cells need to be sorted for further analysis. Therefore, I learned the skill about germ cell sorting in transgenic medaka larvae with fluorescent germ cells. After sorting, certain analysis can be performed, such as microarray. To investigate the contribution of gonadal cell between different sexes and the contribution of different cell types, transplantation must be performed. Thus, I learned the skill of cellular transplantation in medaka embryo. Although more practice is needed, this is a powerful tool in realizing the detail mechanisms during early gonad development in zebrafish.
I would like to thank for NIBB and Prof. Tanaka for granted me this opportunity. And I would also like to thank for all the lab members in Prof. Tanaka’s laboratory. They did help me a lot on experiments and life here in Japan. Without their help, my life here could not go so well. Thank you!

M.G.M Institute of Biosciences and Technology
Mahalunkar Sneha Anand
I have successfully completed the project assigned to me and have got well versed with Medaka fish breeding and fish collection techniques, Micro injection techniques and most of the molecular biology techniques performed in our laboratory. During the internship period I also attended 6th NIBB International Practical Course and 1st NIBB-TLL Joint International Practical Course on “Developmental Genetics of Medaka IV”. This Practical course made me well versed with basic technologies and cutting edge experimental skills which are being used with Medaka including Trangenesis, Sperm Cryo-preservation, screening of the TILLING library, using Infrared laser-evoked gene operator (IR-LEGO) for controlling gene expression and ENU mutagenesis.
I also got an opportunity to attend a two day symposium on Medaka Meet held at National Institute for Basic Biology, Okazaki, Japan.
Apart from my research activity I thoroughly enjoyed my stay at NIBB, Japan. My lab members helped me to learn Japanese language whereas they were also very keen to know about Indian culture which fascinated and touched me a lot. All the members of NIBB were very open and ready to help.
At the last I would like to thank all the members of NIBB to make my stay so comfortable and joyful whereas I would also like to thank Dr. Kiyoshi Sensei for giving me this opportunity. Last and not the least I would like to reveal my wish to do my PhD studies at the same place. I would also like to share some photographs which are life time memories for me which I have attached to the file.

Brandeis University
NAN, Pang
Message from Interns 2010

National Research Center, Egypt
Walaa A Adly
My internship helped me to gain a lot of experiences and skills in a variety of biological fields especially in genome informatics field as well as being able to communicate with different culture and society. My work in the laboratory was very interesting and enthusiastic which we able to combine biological sciences with informational technology in addition to implement scientific programs such as detect genes responsible for diseases by using highly qualified software. The team was very helpful and friendly. Really I liked all people and respected them. Also I would like to thank my professor for his support to me which taught me how I think about things differently.
I am honor and proud of developing my experience and skills of the scientific research in the ideal environment for creativity. Finally I hope to study and complete my PhD. in this innovative and pioneer climate. Now I will summarize my work in genome informatics laboratory.

The University of Heidelberg, Germany
Thomas Auer
The accommodation at the Mishima Lodge was very close to the lab and easily reachable by walking within 10 minutes. All was very well organized and equipped and the room offered all necessary comfort.
Ultimately the time I spend in Okazaki was an unforgettable experience I would not like to miss. Scientifically I was profiting by the expertise of all lab members and learned a lot-besides it was an invaluable introduction into Japanese culture and cuisine. Thank you very much for giving me that opportunity!!!

Allahabad Agricultural Deemed University, India
Nitisha Shrivastava
Whole-mount in situ hybridization (WISH) is a technique widely used to study the expression patterns of developmentally regulated genes. The advent of cell-type specific molecular markers has facilitated the description and analysis of multicellular morphogenesis in many systems. Hybridization of probes prepared from specific cloned genes to whole-mounts can provide three dimensional information, on cells that have accumulated the respective mRNA. Microinjection is one of the most popular techniques used for inserting genetic material in the living organism and studying the overexpression pattern of these genes.

University of Delhi, India
Swati Gupta
Research experience: My stay in Okazaki and research guidance in the laboratory of Prof. Yoshida was remarkable. I couldn’t even imagine of such a unique and innovative work experience in my dreams.
Enormous motivation from Shosei San (Prof. Shosei Yoshida), great support from lab members and enthusiastic surrounding of my friends made my stay the most memorable time of my life. Weekend trips with my lab members and novel experiments in the weekdays always filled me with high energy. Everyday challenges in both laboratory and cooking sessions in night, made me to learn whole lots of thing which are must for the survival of a vegetarian researcher. Accommodation in the Mishima lodge gave me the opportunity to interact with various international students and to learn about their cultural backgrounds. For me this one month period was packed with motivation, enthusiasm, learning, exploring, experimenting, friendship and enjoyment.
My work in the laboratory of Prof. Yoshida was around some genes which were always been favorite between the researchers but still are mysterious about their specific function and expression in the mouse testis. I was lucky enough to get the opportunity to work on these genes i.e. to check their expression in the mouse testis and with the grace of God; within my stay I got the positive signs regarding the expression of these genes. I am highly thankful of Prof. Shosei Yoshida for having faith on me and giving me the opportunity as well as responsibility to work with these genes. Without the help of my lab members i.e. Ichikawa San, Ryo San and Hara san I couldn’t even thought of getting such results and reaching to the inferences about the expression of these genes. Their time and devotion always helped me to think more sincerely and deeply about my work.
Although, for me this one month was a short duration to complete my project but it was a worth time to get the insight of the Japanese culture, research opportunities and for making international friends. I am glad and obliged that NIBB selected me for this internship and gave me the atmosphere to develop scientific skills and research insight.
I once again thank NIBB, especially Prof. Shosei Yoshida and lab members for their professional guidance, cooperation and friendly atmosphere.

VU University Medical Center in Amsterdam, Netherlands
Patricia Faustino
Prof. Nobuyuki Shiina’s team investigates the mechanisms and roles of mRNA transport and local translation in neuronal dendrites using mice. They have identified RNG105, an RNA-binding protein that is a component of RNA granules in dendrites, which is involved in the formation of synapses and neuronal networks. They also have identified several mRNAs associated with RNG105, through microarray analysis. The first part of my project was to clone those mRNAs, which was well succeeded to a big part of them. Then I learned how to do primary neuronal culture from brains of embryonic mice. After this, those cultured neurons were immunostained using antibodies to detect the dendrites and axons. As a result, I could take wonderful pictures of the neurons such as Fig.1 with a confocal fluorescence microscope.
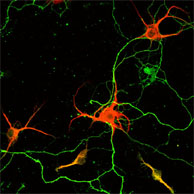
Fig.1
The last part of my project was learning how to transfect neuron cells by calcium-phosphate method. We transfected the neurons with Na+/K+ ATPase subunit isoforms, that were previously identified as RNG105-bound mRNAs. We could then visualize their localization in the neurons and live imaging after activating translation.
I am deeply grateful for the opportunity this internship program provided me. I had the chance of learning exciting techniques with great researchers in a fantastic environment. I have to thank to Prof. Shiina for hosting me in his lab, to Nakayama-san for all the patience in teaching me, (specially in dissecting brains of embryonic mice, which was not easy) and also to Matsuda-san for all the help in the lab. I met several interesting persons that allowed me to experience the wonderful Japanese hospitality and gave me great insights of the Japanese culture.
Report List
Message from Interns 2024
Message from Interns 2023
Message from Interns 2022
Message from Interns 2021
Message from Interns 2020
Message from Interns 2019
Message from Interns 2018
Message from Interns 2017
Message from Interns 2016
Message from Interns 2015
Message from Interns 2014
Message from Interns 2013
Message from Interns 2012
Message from Interns 2011
Message from Interns 2010














