12.1 Constitutive overexpression of cDNA or cDNA
with tags
Tomomichi Fujita
CaMV 35S promoter is a weak promoter in P.
patens, although it gives
enough resistance for drug selection with resistant genes. The rice actin
promoter and the CaMV35S derived 7113 promoter are stronger promoters than CaMV
35S. As platform sites for targeting an overexpression construct, PpMADS2,
Pphb7, and BS213 loci are used (Pphb7, Sakakibara et al. 2003; BS213, Schaefer
et al. 1997).
Caution
(1) When Pphb7 is disrupted, development of
rhizoid becomes abnormal. Development of protonemata is not distinguished from
that in wild type (Sakakibara et al. 2003). We do not find any phenotypic
effects by disruptions of PpMADS2 and BS213.
(2) Gene silencing is sometimes observed ccurs
in some lines with overexpression constructs. The gene silencing is transmitted
from generation to generation, although gene expression is recovered in the
mucilage hairs of a gametophore and the foot of a sporophyte. If you select a
line with a single insertion, gene silencing is rare. On the other hand, gene
silencing usually happens for multi copy insertion lines. Northern or western
analyses are recommended to confirm whether the targeted gene is effectively
overexpressed. It is a good idea to fuse the targeted gene with GFP, although
it is necessary to confirm the fusion protein is functional.
(3) Actin promoter works in protonemata
well but much weaker in gametophores. The 7113 promoter is recommended to
overdrive in gametophores.
(4) 7113 promoter was kindly provided from
Dr. I. Mitsuhara and Dr. Ohashi (Mitsuhara et al. (1996), Plant Cell Physiol.
37: 49-59). Actin promoter from rice
was kindly provided from Dr. Wu (Zhang et al. (1991), Plant Cell, 3:1155-1165).
Rice actin promoter worked >10 times stronger than 7113 promoter in P.
patens protoplasts (Fujita
et al. unpublished).
Available plasmids (I)
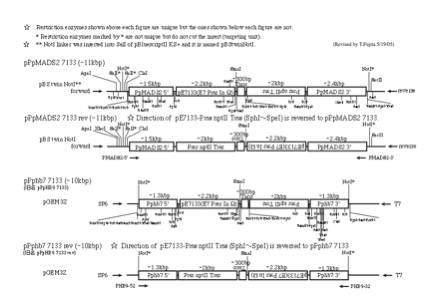
- E. coli harboring a plasmid with the 7113 promoter should be
cultured at 30ºC rather than 37ºC.
Truncation of 7113 promoter sequence is sometimes found at 37ºC.
- NotI is usually used to linearize the plasmid for
transformation.
- To screen stable transformants, we
perform green PCR (See a section of Green PCR for PCR strategy).
Available plasmids (II)
(1)
PpMADS2 target, rice Actin promoter
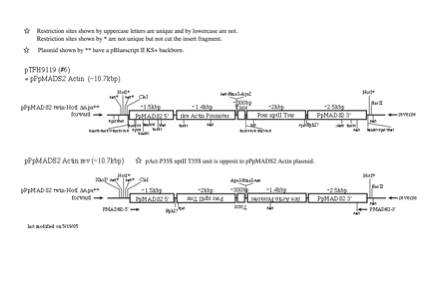
NotI site is usually used to linearize the plasmid for transformation.
(2) Pphb7 target, rice actin promoter
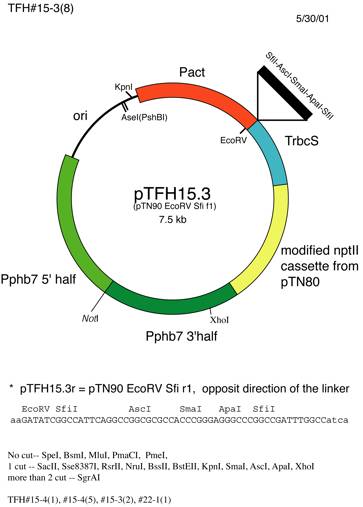
NotI site is usually used to linearize the plasmid for transformation.
(3) BS213 target,
7113 promoter, gateway construction, with tag sequence (His, mRFP, citrine)
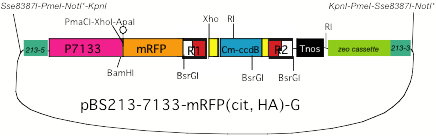
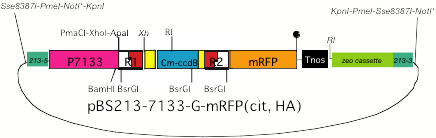
*As LR reaction with pENTR Directional TOPO
brings NotI site into
the resultant product, NotI
cannot be used for excision of a target fragment.