11. RNA interference
Yuji Hiwatashi
Introduction
RNA interference (RNAi) takes advantage of
the unique ability of double-stranded RNA (dsRNA) molecules to induce
posttranscriptional gene silencing in a highly specific manner. A RNAi
construct is introduced to protoplasts by PEG-mediated transformation. RNAi
becomes effective at 2 to 3 cell stage after protoplast regeneration. Loss of function phenotype of FtsZ1, the
larger chloroplast than in wild type, is observed only in an apical cell at
three or more cell protonemata 6 days after transformation. RNAi for GFP is
effective 4 days after transformation, and GFP signals were diminished in all
cells.
Steps for RNAi experiments:
1) Clone a DNA fragment of a target gene into entry
vector pENTR/D-TOPO
2) Transfer the DNA fragment in pENTR/D-TOPO to a
destination vector pPI1 using the GATEWAY system.
3) Co-transformation of protoplasts with both the pPI1
derived-RNAi plasmid and a plasmid harboring a visual expression marker.
4) Observation of regenerated protoplasts expressing the
visual marker gene.
Materials
・ pENTR Directional TOPO Cloning Kit (Invitrogen)
・ Wizard Mini-Prep kit (Promega)
・ Hi Speed Midi kit (QIAGEN)
・ Reagents for PCR amplification
・ Reagents for agarose gel electrophoresis
・ Gateway LR Clonase Enzyme Mix (Invitrogen)
・ QIAquick gel extraction kit (QIAGEN)
・ Reagents for PEG-mediated transformation
・ pPI plasmid (see fig. 1). This gateway plasmid for RNAi was originally designed and constructed by Dr. Tsuyoshi Nakagawa (Shimane Univ., Japan) and modified by Y. Hiwatashi.
Procedure
1. Cloning of a gene of interest into the entry vector
pENTR/D-TOPO.
Refer a manual of pENTR Directional TOPO
Cloning Kit (cat.no.K2400-20).
1) Design your PCR primer with CACC at the 5’ end
(modification of the 3’ primer is not necessary). Amplify your gene with a proofreading enzyme (see below).
PCR total 20 µl
1
ul template cDNA
2
ul 10x KOD+ buffer
2
ul 2.5 mM dNTPs
0.8
ul 25 mM MgSO4
0.6
ul 10 pmol/ul primer1
0.6
ul 10 pmol/ul primer2
0.4
ul 1 u/ul KOD+
12.6
ul H2O
PCR cycle
94C,
2 min
94˚C,
15 s ━━┓
Tm-5˚C,
30 s x 30
68˚C,
1 m/1kb ━━┛
4˚C
2) Mix your PCR product with a pENTR vector, transform
E.coli, and plate.
TOPO reaction total
1.5 µl
0.25
ul Salt
0.5
ul H2O
0.25
ul pENTR/D-TOPO
0.5
ul PCR product (5-20 mg)
↓
Incubate
at room temperature for more than 5 min.
↓
Transform
a competent cell XL10-GOLD with all solution, then spread all cells on a LB
plate supplemented with 50 mg/l kanamycin, and incubate at 37˚C overnight.
3) Select positive clones by colony PCR.
4) Mini-prep a positive plasmid with Wizard Mini prep Kit
or equivalents.
5) Digest the plasmid with appropriate restriction
enzyme.
Restriction enzyme
treatment total 10 µl
3
ul Plasmid
1
ul 10 x H
1
ul 0.1% BSA
0.5
ul 10 u/µl EcoRV
0.5
ul 10 u/µl NotI
4
ul H2O
↓
Incubate at 37˚C for 1
hrs.
↓
Perform gel-electrophoresis.
6) Confirm the direction and sequence of the fragment by
sequencing
2. Transfer the DNA fragment cloned in pENTR/D-TOPO into RNAi
vector pPI1 (Figure 1).
Refer to a manual of Gateway LR Clonase Enzyme
Mix (cat.no.11791-019).
1) Digest the pPI1 vector with a restriction enzyme to be
a linear form .
1.5
ul 1 µg/µl pPI1
1
ul 10 x H
0.5
ul 10 u/µl Xho I
8
ul H2O
↓
Incubate at 37˚C for
more than 1 hrs.
↓
Perform
gel-electrophoresis.
↓
Recover the digested plasmid
with QIAquick Gel extraction kit.
2) Perform a LR reaction.
1
ul Entry clone (supercoiled, ~60 ng)
1
ul ~ 60 ng/µl linearized pPI1
0.8
ul 5x LR Clonase Rxn. Buffer
0.4
ul TE
↓
Add 0.8 µl of LR
clonase enzyme mix, and then mix by pipetting
↓
Incubate at 25˚C for
more than 2 hrs.
↓
Add 0.4 µl of
proteinase K, and then mix by pipetting.
Incubate at 37˚C for 10 min.
3) Transform a competent cell DH10B (or DH5alpha) with the
product in the previous step. Spread all
cells on a LB+Amp plate and incubate 30˚C over night.
4) Select a correct RNAi vector by colony PCR. Since the LR reaction allows inverted
direction of an intron in the RNAi vector, confirm its direction by PCR. Confirm an integration of the fragment,
followed by the direction of the intron.
First colony PCR(total 20µl)
colony
inoculation
2
ul 10xEx Taq
2
ul 2.5 mM dNTPs
0.5
ul 10 pmol/ul 35S mini-F (CTAATCTTCGCAAGACCCTTCCTC)
0.5
ul 10 pmol/ul antisense primer
0.125
ul 5 u/ul Ex Taq
14.875
ul H2O
PCR cycle
94˚C,
30 s ━━┓
58˚C,
30 s x 30
72˚C, 1 m/1 kb ━━┛
4˚C
Second colony PCR (total 20 µl)
2
ul 10xEx Taq
2
ul 2.5 mM dNTPs
0.5
ul 10 pmol/ul 35S mini-F
0.5
ul 10 pmol/ul GFP1r1KpnI or GFPf1SpeI
0.125
ul 5 u/ul Ex Tag
14.875
ul H2O
GPA1r1KpnI (ACCGGTACCTGCATATAACCTGC)
GPAf1SpeI (GATACTAGTGGTCGGTAACGGTCGG)
PCR cycle
94˚C,
30 s ━━┓
58˚C,
30 s x 25
72˚C, 1 m ━━┛
4˚C
The amount of PCR amplified
fragment with 35S mini-F and GPA1r1KpnI primers should be more than that with 35S
mini-F and GPAf1SpeI primers on agarose gel electrophoresis in candidate
clones.
5) Mini-prep a positive clone with Wizard Mini Prep Kit
(Promega) or equivalents.
6) Digest the plasmid with appropriate
restriction enzymes.
2
ul plasmid
1
ul 10 x M 10x M
0.5
ul 10 u/µl Xho I 10 u/µl XhoI
0.5
ul 10 u/µl Sac I 10 u/µl KpnI
6
ul H2O
↓
Incubate at 37˚C for
more than 1 hrs.
↓
Perform gel-electrophoresis.
7) Sequence the plasmid with the 35S mini-F and reverse
primers.
8) Prep the plasmid in a large-scale with QIAGEN hi Speed
Midi Kit (QIAGEN).
3. Introduction of RNAi vector to a protoplast.
Protoplasts are co-transformed with the
constructed RNAi plasmid and a visual selection marker plasmid since the pPI1
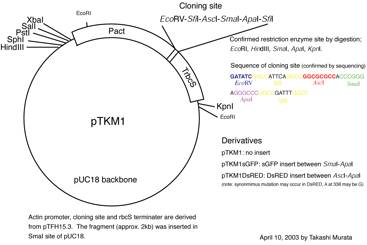
itself have no marker cassette. For a visual maker plasmid, pTKM-GFP, pTKM-RFP,
and pTKM-DsRED2 are used.
1) Prepare a marker plasmid.
2) Co-transform protoplasts with a mixture of 10 µg
of the RNAi plasmid and 10 µg of a marker plasmid. As a control,
co-transform with the original pPI1 plasmid instead of the RNAi plasmid.
3) Incubate the transformed protoplasts.
4. Observation
Regenerated protonemata expressing a marker
gene such as GFP and mRFP are observed.
Figure 1. pPI1
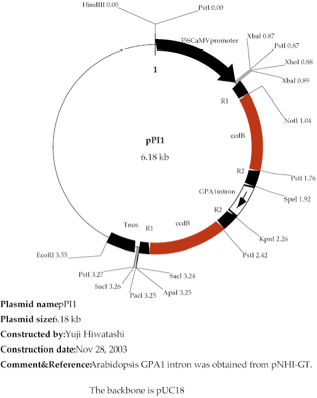
Figure 2. pTKM-GFP