| DEPARTMENT OF CELL BIOLOGY | ||||
|
||||
Higher plant cells contain several distinct organelles
that play vital roles in cellular physiology. During proliferation
and differentiation of the cells, the organelles often undergo dynamic
changes. The biogenesis of new organelles may occur, existing organelles
may undergo a transformation of function, while other organelles may
degenerate. Because the dynamic transformation of organellar function
(differentiation of organelles) is responsible for flexibility of
differentiation events in higher plant cells, the elucidation of regulatory
mechanisms underlying organelle transformation are currently studied
in this division. Dramatic metabolic changes which underlie the shift from heterotrophic
to autotrophic growth occur in greening of seed germination. Accompanying
these metabolic changes, many constitutive organelles are functionally
transformed. Etioplasts differentiate into chloroplasts and mitochondria
acquire the ability to oxidize glycine. Glyoxysomes which are microbodies
engaged in the degradation of reserve oil via b-oxidation
and the glyoxylate cycle, are transformed into leaf peroxisomes that
function in several crucial steps of photorespiration. After the functional
transition of glyoxysomes to leaf peroxisomes during the greening
of pumpkin cotyledons, the reverse transition of leaf peroxisomes
to glyoxysomes occurs during senescence. The functional transformation
between glyoxysomes and leaf peroxisomes is controlled by gene expression,
alternative splicing, protein translocation and protein deg-radation.
We now engage in proteomic and transcriptomic analyses of the reversible
peroxisomal transition in Arabidopsis cotyledons. Enzymes localized in plant peroxisomes are synthesized in the cytosol, and function after their post-translational transport into peroxisomes. Almost all of the peroxisomal matrix proteins are known to contain one of two targeting signals (PTS1 and PTS2) within the molecules. PTS1 is a unique tripeptide sequence found in carboxyl terminus of the mature proteins. The permissible combinations of amino acids for PTS1 in plant cells are [C/A/S/P]-[K/R]-[I/L/M]. In contrast, PTS2 is involved in a cleavable amino terminal presequence of peroxisomal proteins that are synthesized as precursor protein with larger molecular mass. PTS2 consists of a consensus sequence [R]-[L/Q/I]-X5-[H]-[L]. We identified 256 gene candidates of PTS1- and PTS2-containing proteins
and other 30 genes of non-PTS-containing proteins from Arabidopsis genome. Custom-made DNA microarray covering all these genes was used
to investigate expression profiles of the peroxisomal genes in various
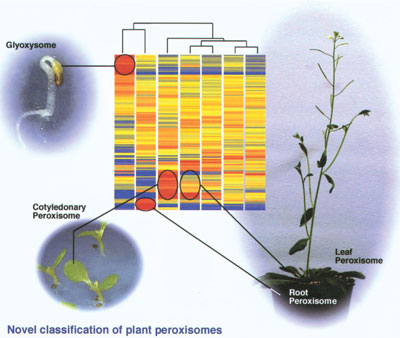
organs. Statistical analyses revealed that the peroxisomal genes could
be divided into five groups in terms of their transcription. One group
showed ubiquitous expression in all organs examined, while the other
four were classified as showing organ-specific expression in seedlings,
cotyledons, roots and in both cotyledons and leaves. These data proposed
more detailed description of differentiation of plant peroxisomes
(Fig. 1).
In parallel, we made two-dimensional protein map of glyoxysomes and
leaf peroxisomes isolated from Arabidopsis. Peptide MS fingerprinting
analyses allowed us to identify novel proteins exists in either glyoxysomes
or leaf peroxisomes. Some of these proteins contain no obvious PTS1
and PTS2. Of these, we characterized GPK1 as a novel protein kinase
in glyoxysomes. To better understand peroxisome biogenesis, we mutagenized seeds of
transgenic Arabidopsis, GFP-PTS1, in which peroxisomes with
normal size and number can be visualized with GFP, and screened a
number of Arabidopsis mutants with aberrant peroxisome
morphology (apm mutants) based on the different pattern
of GFP. The apm mutants were classified into four classes. These were
mutants with (1) long peroxisomes, (2) giant peroxisomes, (3) GFP
fluorescence in the cytosol as well as in peroxisomes, and (4) other
distributions of GFP.
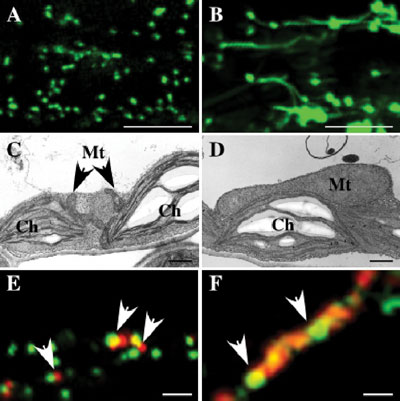
In one of these mutants, apm1, the peroxisomes are long
and reduced in number, apparently as a result of inhibition of division
(Fig. 2). APM1 gene encodes DRP3A (Dynamin related protein 3A). Interestingly, mutations in APM1/ADL2A also
caused aberrant morphology of mitochondria. The growth of Arabidopsis,
which requires the cooperation of various organelles including peroxisomes
and mitochondria, is repressed in apm1, indicating that
the changes of morphology of peroxisomes and mitochondria reduce
the efficiency of metabolism in these organelles. These findings
indicate that the same dynamin molecule is involved in peroxisomal
and mitochondrial division in higher plants. Precursor-accumulating (PAC) vesicle, mediates the transport of storage protein precursors from endoplasmic reticulum (ER) to protein storage vacuoles in maturing pumpkin seeds. PAC vesicles had diameters of 300 to 400 nm, are derived from ER and contained an electron-dense core of storage proteins. PV72, a type I integral membrane protein was found on the membrane of the PAC vesicles. PV72 has been shown to bind to the precursor of pumpkin 2S albumin, implying that PV72 functions as a vacuolar sorting receptor (VSR) for storage proteins in pumpkin seeds. Arabidopsis has seven homologue of PV72. The homologue closely related to PV72 was designated AtVSR1. We used a reverse-genetic approach to explore the function of AtVSR1. Two T-DNA insertion mutants (atvsr1-1 and atvsr1-2) missort storage proteins by secreting them from cells, and abnormally accumulate the precursors of storage proteins, together with the mature forms of these proteins in the seeds. These finding demonstrate a receptor-mediated transport of seed storage proteins to protein storage vacuoles in higher plants. ER bodies are another ER-derived compartment specific to the Brassicaceae,
including Arabidopsis. ER bodies are rod-shaped structures
(5 µm long and 0.5 µm wide) that is surrounded by ribosomes.
ER bodies can be visualized in transgenic plants of Arabidopsis (GFP-h) expressing green fluorescent protein fused with
an ER retention signal (GFP-HDEL). ER bodies were widely distributed
in the epidermal cells of whole seedlings. In contrast, rosette
leaves had no ER bodies. nai1 is an Arabidopsis mutant in which ER bodies were hardly detected in whole plants.
Analysis of the nai1 mutant reveales that a b-glucosidase
with an ER-retention signal (KDEL), called PYK10, is the main component
of ER bodies. The putative biological function of PYK10 and the
inducibility of ER bodies in rosette leaves by wound stress suggest
that the ER body functions in the defense against herbivores. Vacuolar processing enzyme (VPE) belongs to the cysteine protease family C13. This family is found in various eukaryote organisms including higher plants and animals. VPE was originally identified as an enzyme responsible for the processing and maturation of seed storage proteins in plants. VPE exhibit substrate specificity toward an asparagine residue, the amino acid well conserved at the P1 position in the processing sites of various vacuolar/lysosomal proteins. Plant VPE homologues were separated to two subfamilies: one seed type and the other vegetative type. We have identified one seed type (bVPE) and two vegetative type (aVPE and gVPE) VPE genes from Arabidopsis. We isolated six Arabidopsis mutants that accumulate detectable amounts of the precursors of the storage proteins. All mutants had a defect in the bVPE gene, indicating that bVPE is involved in the processing of storage proteins in vivo. We further generated various mutants lacking different VPE isoforms: aVPE, bVPE and/or gVPE. avpe-1/bvpe-3/gvpe-1 triple mutant seeds accumulate no properly processed mature storage proteins. Instead, large amounts of storage protein precursors are found in the seeds of this mutant. In contrast to bvpe-3 seeds, which accumulate both precursors and mature storage proteins, the other single (avpe-1 and gvpe-1) and double (avpe-1/gvpe-1) mutants accumulate no precursors in their seeds at all. Therefore, the vegetative type VPEs, aVPE and gVPE, compensates for the deficiency in bVPE in bvpe mutant seeds. To explore the physiological function of VPE in mammals, we generated
and characterized VPE-deficient mice. VPE was abundantly expressed
in kidney and localized in the late endosomes of the proximal tuble
cells. Disruption of the VPE gene led to the enlargement
of lysosomes in these cells in an age-dependent manner, which suggests
that the materials to be degraded are being accumulated within the
lysosomal compartments. The processing of the lysosomal proteases,
cathepsins B, H, and L, from the single-chain forms into the two-chain
forms was completely defected in the deficient mice. Thus, the VPE
deficiency caused the accumulation of macromolecules in the lysosomes,
highlighting a pivotal role of VPE in the endosomeal/lysosomal degradation
system. Molecular chaperones are cellular proteins that function in the
folding and assembly of certain other polypeptides into oligomeric
structures but that are not, themselves, components of the final
oligomeric structure. To clarify the roles of molecular chaperones
on cell differentiation, we have purified and characterized chaperonin
and Hsp70s and analyzed their roles in the translocation of proteins
into chloroplasts. |
||||
Publication List: Fukao, Y., Hayashi, M., Hara-Nishimura, I. and Nishimura, M. (2003) Novel glyoxysomal protein kinase, GPK1, identified by proteomic analysis of glyoxysomes in etiolated cotyledons of Arabidopsis thaliana. Plant Cell Physiol. 44, 1002-1012. Hayashi, M. and Nishimura, M. (2003) Entering a new era of research on plant peroxisomes. Cur, Opi. Plant Sci. 6, 577-582. Kamada, T., Nito, K., Hayashi, H., Mano, S., Hayashi, M. and Nishimura, M. (2003) Functional differentiation of peroxisomes revealed by expression profiles of peroxisomal genes in Arabidopsis thaliana. Plant Cell Physiol. in press Kamigaki, A., Mano, S., Terauchi, K., Nishi, Y., Tachibe-Kinoshita, Y., Kondo, M., Nito, K., Hayashi, M., Nishimura, M. and Esaka, M. (2003) Identification of peroxisomal targeting signal of pumpkin catalase and the binding analysis with PTS1 receptor. Plant J. 33, 161-175. Kurisu, M., Morita, M., Kashiwayama, Y., Yokota, S., Hayashi, H., Sakai, Y., Ohkuma, S., Nishimura, M. and Imanaka, T. (2003) Existence of catalase-less peroxisomes in Sf21 insect cells. Biochem. Biophys. Res. Commun. 306, 169-176. Matsushima, R., Hayashi, Y., Yamada, K., Shimada, T., Nishimura, M. and Hara-Nishimura, I. (2003) The ER body, a novel endoplasmic reticulum-derived structure in Arabidopsis. Plant Cell Physiol. 47, 661-666. Okamato, T., Shimada, T., Hara-Nishimura, I., Nishimura, M. and Minamikawa, T. (2003) C-terminal KDEL sequence of a KDEL-tailed cysteine protease (sulfhydryl-endopeptidase) is involved in formation of KEDL vesicle and in efficient vacuolar transport of sulfhydryl-endopeptidase. Plant Physiol. 132, 1892-1900. Ono, K., Kondo, M., Osafune, T., Miyatake, K., Inui, H., Kitaoka, S., Nishimura, M. and Nakano, Y. (2003) Presence of glyoxylate cycle enzymes in the mitochondria of Euglena gracilis. J. Eukaryo. Microbiol. 50, 92-96. Shimada, T., Fuji, K., Kondo, M., Nishimura, M. and Hara-Nishimura, I. (2003) A vacuolar sorting receptor for seed storage proteins in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 100, 16095-16100. Shimada, T., Yamada, K., Kataoka, M., Nakaune, S., Koumoto, Y., Kuroyanagi, M., Tabata, S., Kato, T., Shinozaki, K., Seki, M., Kobayashi, M., Kondo, M., Nishimura, M. and Hara-Nishimura, I. (2003) Vacuolar processing enzymes are required for proper processing of seed storage proteins in Arabidopsis thaliana. J. Biol. Chem. 278, 32292-32299. Shirahama-Noda, K., Yamamoto, A., Sugihara, K., Hashimoto, N., Asano, M., Nishimura, M. and Hara-Nishimura, I. (2003) Biosynthetic processing of cathepsins and lysosomal degradation in asparaginyl endopeptidase-deficient mice. J. Biol. Chem. 278, 33194-33199 Tamura, K., Shimada, T., Ono, E., Tanaka, Y., Nagatani, A., Higashi, S., Watanabe, M., Nishimura, M. and Hara-Nishimura I. (2003) Why green fluorescent fusion proteins have not been observed in the vacuoles of higher plant? Plant J. 35, 545-555. Watanabe, E., Shimada, T., Tamura, K., Matsushima, R., Koumoto, Y., Nishimura, M. and Hara-Nishimura, I. (2003) An ER-localized form of PV72, a seed-specific vacuolar sorting receptor, interferes the transport of an NPIR-containing proteinase in Arabidopsis leaves. Plant Cell Physiol. in press Yamada, K., Nishimura, M. and Hara-Nishimura, I. (2003) The slow wound-response of gVPE is regulated by endogenous salicylic acid in Arabidopsis. Planta in press Yamamoto, Y., Nishimura, M., Hara-Nishimura, I. and Noguchi, T. (2003) Behavior of vacuoles during pollen development and maturation in Arabidopsis thaliana. Plant Cell Physiol. 44, 1192-1201. |