| DIVISION OF SPECIATION MECHANISMS I | ||||
|
||||
| 1) from Utah State University, USA |
||||
| All living organisms evolved from a
common ancestor that lived more than 3.5 billion years ago, and the
accumulation of mutations in their genomes has resulted in the present
biodiversity. Traces of the evolutionary process are found in the
genomes of extant organisms. By comparing the gene sequences and gene
networks of different organisms, we can infer (1) the phylogenetic
relationships of extant organisms and (2) the genetic changes that
caused the evolution of morphology and development. The inferred phylogenetic
relationships provide important insights into problems in various
fields of evolutionary biology. Our group focuses on biogeography,
the evolution of morphological traits, and systematics in a wide range
of taxa. Concerning the evolution of morphology and development, we
hope to explore the genetic changes that led to the evolution of the
plant body plan. We have selected Arabidopsis (angiosperm), Gnetum
(gymnosperm), Ginkgo (gymnosperm), Ceratopteris (pteridophyte), Physcomitrella
(bryophyte), and some green algae as models to compare the functions
of genes involved in the development of both reproductive and vegetative
organs in land plants. The first green alga cell evolved via symbiosis between an ancestral
non-photosynthetic eukaryote and a cyanobacterium. Cyanobacteria now
exist as chloroplasts in the host cell. The factors and mechanisms
of chloroplast movement are being investigated to reveal the molecular
mechanisms used to "domesticate" cyanobacteria as organelles.
Analyses of cytosolic calcium icon concentration and cytoskeleton
organization during chloroplast movement in the moss Physcomitrella
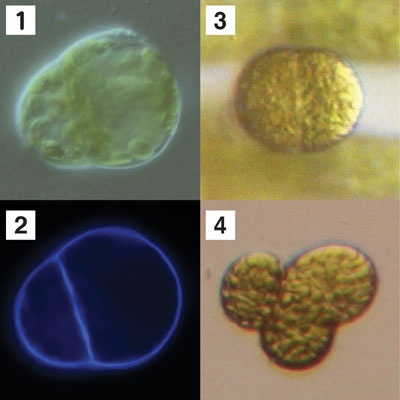
patens is in progress by a team directed by Y. Sato. The first evolutionary step from unicellular to multicellular organisms
is to form two different cells from a single cell via asymmetric cell
division. The first cell division of a protoplast isolated from the
protonemata of the moss Physcomitrella patens is asymmetric
regarding to its shape and nature, and gives rise to an apical meristematic
cell and a differentiated non-meristematic cell. A systematic overexpression
screening for genes involved in asymmetric cell division of protoplasts
in P. patens is in progress by a team directed by T. Fujita.
We constructed three full-length cDNA libraries from non-treated,
auxin-treated, and cytokinin-treated protonemal cells of P. patens,
then determined the sequences of more than 40,000 cDNAs from the both
ends (Nishiyama, Fujita et al. 2003). We used these clones as materials
for the overexpression screening. Individual cDNAs were selected based
on their sequence, subcloned under a constitutive promoter and introduced
into the protoplasts of P. patens for transient expression.
We observed and categorized phenotypes of the regenerating protoplasts.
Thus far we identified many cDNAs, whose overexpression resulted in
symmetric cell division rather than asymmetric cell division, isotropic
outgrowth with no polarity or curved cells showing incorrect direction
of growth. These preliminary results indicate that some of these genes
likely function for polarity formation and/or asymmetric cell division
of the protoplasts. Functional analyses of these genes should help
us to understand molecular mechanisms of how plants generate distinct
cell lineages to build their multicellular bodies.
The most prominent difference between plant and animal cells is that
plant cells have a cell wall and do not move during development. Therefore,
the plane of cell division and the direction of cell elongation, which
are regulated by cortical microtubules, determine the morphology of
differentiated tissues and organs. g-tubulin is a protein that is essential
for the formation of microtubules in animal cells. We found that g-tubulin
is located at the end of cortical microtubules, and a loss of g-tubulin
due to gene silencing causes a malformed organ with irregularly shaped
cells. In vitro experiments using isolated plasma membrane/microtubule
complexes suggested that g-tubulin attaches
onto the side of existing cortical microtubules, and initiates a new
cortical microtubule from it. The nucleus is known to initiate microtubules
after cell division. We hypothesize that microtubules formed around
the nucleus elongate to cell surface, and trigger initiation of cortical
microtubules via attachment of g-tubulin.
Once cortical microtubules are formed, they can turnover without microtubules
from the nucleus. Factor(s) responsible for attachment of g-tubulin
onto the side of microtubules is a key element responsible for the
difference between plant and animal cells. Isolation of the factor(s)
responsible for attachment of g-tubulin
onto the side of microtubules by biochemical and other approaches
is inprogress by a team directed by T. Murata. Cells of the moss P. patens gametophytes are an excellent
model to study dynamics of cytoskeleton because of the easiness for
observation and the feasibility of gene-targeting. Transformants with
reporter constructs by fusing a GFP with P. patens a-tubulin
or an actin binding domain of mouse talin were established by Y. Sato
and collaborators to visualize microtubules and actin filaments. These
transformants will be useful to investigate the dynamics of microtubules
and actin filaments in the processes of cell division, cell elongation,
and chloroplast movement. Postembryonic growth of land plants occurs from the meristem, a localized region that gives rise to all adult structures. Meristems control the continuous development of plant organs by balancing the maintenance and proliferation of stem cells, and directing their differentiation. Meristem initiation and maintenance is a fundamental question in plant development research. However, the molecular mechanisms involved in meristem initiation and maintenance have not been studied in detail because most loss-of-function mutants are lethal. In the moss Physcomitrella patens, the developmental process of meristem is well defined at the cellular level, and gene targeting based on homologous recombination is feasible. Thus, meristem development in P. patens is used as a model system for studies of meristem development in land plants. We established approximately 20,000 gene- or enhancer-trap lines in P. patens to clone genes involved in meristem development. Seven lines, exhibiting reporter gene (uidA) expression preferentially in the apical cells, were isolated. Corresponding genes in three of these trap lines were identified as encoding kinesin- and ubiquitin-like proteins, and an unknown protein. Disruption of the gene encoding ubiquitin-like protein suggests that the gene be involved in cell division and elongation through microtubule organization. The functions of other genes in the meristem are currently under investigation by a team directed by Y. Hiwatashi. The morphology of the shoot meristem in land plants varies. To investigate
whether the molecular mechanisms of shoot development in angiosperms
are conserved in other land plants, the functions of the KNOX homeobox,
and the ZWILLE, NAC, and PIN genes, which are indispensable for shoot
meristem development in angiosperms, are being studied in the fern
Ceratopteris and the moss Physcomitrella. Rhizoids are multicellular filaments with similar functions to root
hairs, and are observed in a wide range of plants. A team directed
by K. Sakakibara, who was a graduate student in our lab, examined
mechanisms underlying rhizoid development in the moss, Physcomitrella
patens (Sakakinara et al. 2003).
The flower is the reproductive organ in angiosperms, and floral homeotic
genes, such as MADS-box genes and FLO/LEAFY genes, regulate floral
organ identity. To investigate the origin of floral homeotic genes,
the functions of MADS-box genes and the FLO/LEAFY genes in gymnosperms
(Gnetum, Ginkgo, and cycads), a fern (Ceratopteris),
a moss (Physcomitrella), and three green algae (Chara,
Coleochaete, and Closterium) are being analyzed. The mosses and flowering plants diverged more than 400 million years
ago. The mosses have haploid-dominant life cycles, while the flowering
plants are diploid-dominant. The common ancestors of land plants are
inferred to have been haploid-dominant, suggesting that genes used
in the diploid body of flowering plants were recruited from the genes
used in the haploid body of their ancestors during the evolution of
land plants. Reproductive isolation is the first step in speciation. To obtain insight into reproductive isolation, several receptors specifically expressed in the pollen tube are being studied to screen for the receptors that are involved in pollen tube guidance by a team directed by S. Miyazaki. Polyploidization is a major mode of speciation in plants, although
the changes that occur after genome duplication are not well known.
Polyploid species are usually larger than diploids, but the mechanisms
responsible for the size difference are unknown. To investigate these
mechanisms, tetraploid Arabidopsis was established and its gene expression
patterns are being compared to those of diploid wild-type plants using
microarrays. We determined the complete nucleotide sequence of the chloroplast
genome of the leptosporangiate fern, Adiantum capillus-veneris L. (Pteridaceae) as a collaboration work with P. Wolf and C. Rowe
from Utah State University (Wolf et al. 2003). Phylogenetic analysis
of basal land plants using the sequence is in progress by T. Nishiyama.
Many unusual start codons and internal stop codons were found, suggesting
that extensive number of RNA editing exists in the genome. |
||||
Publication List: Kofuji, R., Sumikawa, N., Yamasaki, M., Kondo, K., Ueda, K., Ito, M. and Hasebe, M. 2003. Evolution and divergence of MADS-box gene family based on genome wide expression analyses. Mol. Biol. Evol. 20: 1963-1977. Sakakibara, K., Nishiyama, T., Sumikawa, N., Kofuji, R., Murata, T. and Hasebe, M. 2003. Involvement of auxin and a homeodomain-leucine zipper I gene in rhizoid development of the moss Physcomitrella patens. Development 130: 4835-4846. Nishiyama, T., Fujita, T., Shin-I, T., Seki, M., Nishide, H., Uchiyama, I., Kamiya, A., Carninci, P., Hayashizaki, Y., Shinozaki, K., Kohara, Y., and Hasebe, M. 2003. Comparative genomics of Physcomitrella patens gemetophytic transcriptome and Arabidopsis thaliana: Implication for land plant evolution. Proc. Natl. Acad. Sci. USA 100: 8007-8012. Wolf, P.G., Rowe, C.A., Sinclair, R.B., and Hasebe, M. 2003. Complete nucleotide sequence of the chloroplast genome from a leptosporangiate fern, Adiantum capillus-veneris L. DNA Res. 10: 59-65. Tanabe, Y., Uchida, M. Hasebe, M., and Ito, M. 2003. Characterization of the Selaginella remotifolia MADS-box gene. J. Plant Res. 116: 71-75. Itoh, Y., Hasebe, M., Davies, E., Takeda, J., and Ozeki, Y. 2003. Survival of Tdc transposable elements of the En/Spm superfamily in the carrot genome. Mol. Gen. Genomics 269: 49-59. Rivadavia, F., Kondo, K., Kato, M., and Hasebe, M. 2003. Phylogeny of the sundews, Drosera (Droseraceae) based on chloroplast rbcL and nuclear 18S ribosomal DNA sequences. Amer. J. Bot. 90: 123-130. |