| DIVISION OF SPECIATION MECHANISMS I | |||||||
|
|||||||
| Our research is focusing to understand
mechanisms underlying memory, formation and evolution of the brain.
For one approach to understand these questions, we are studying the
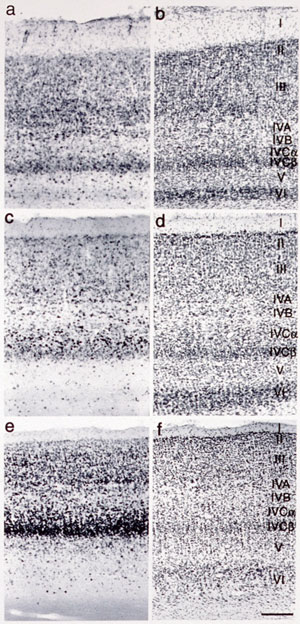
genes that are expressed in specific areas of the primate neocortex.
Using differential display method, we obtained three genes that showed
marked differences within areas of the primate neocortex. Our second
approach is to understand informational processing in the brain underlying
learning behaviors by examining gene expression. The third approach
is to study voles of CNTF in the nervous system. The neocortex, most remarkably evolved in primates, plays the major
role in higher functions of the brain. It is known to be divided into
distinct functional and anatomical areas and has been a matter of
debate what extent the area of the neocortex are genetically and environmentally
deter-mined. It is also puzzling why, during the evolution of mammals,
the neocortex was most markedly expanded while the number of the genes
in the mammal was little changed. To access these questions, we studied
gene expression within different areas of the neocortex. In summary, our studies thus far revealed the following points.
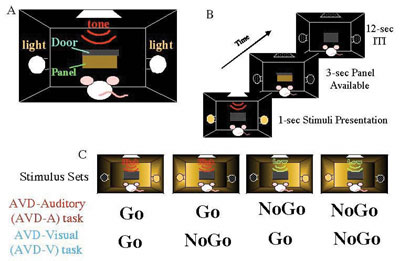
In order to study informational processing underlying the declarative and non-declarative memory at molecular and cellular levels in the brain, we employed c-Fos mapping techniques, for which we used gene expression of c-Fos. There have been an increasing number of studies using c-Fos as markers to examine neuronal activities ever since c-Fos induction by electrical stimulation was found by Morgan and Curran. However, many sensory stimuli per se are now known to cause c-Fos induction. So, we should be very careful to distinguish the c-Fos expression that is caused by learning process from that caused by sensory stimuli. For this purpose, it is necessary to use behavioral systems that are able to distinguish the two. Although a few behavioral systems in rodents have been successfully used for physiology, animal behavior and recently for analyses of knockout mice, little behavioral systems in fact distinguish it. Therefore, we prepared ourselves for using two behavioral systems, which represent declarative and non-declarative memory, respectively. In collaboration with professor Yoshio Sakurai (Kyoto University)
who developed audio-visual discrimination task (AVD-task). In this
task, a rat was asked to choose either an audio cue (a high tone or
low tone) or a visual cue (a light from the right or the left) to
obtain a food pellet . We found that the visual and audio tasks enhanced
the specific expression of c-Fos in the visual and audio cortices,
respectively. Among the early visual and auditory pathways examined,
c-Fos was specifically induced in the cortices but not in the earlier
pathways, suggesting the neural modulation of the neocortex depending
on the types of the tasks. Interestingly, the task-dependent Fos expression
was only observed in excitatory neurons in the relevant sensory cortices.
Although this AVD task system is quite powerful to analyze a kind of task above described and presumably very useful for studying underlying molecular and cellular mechanisms because of advantages using rodents, one problem is that the auditory stimuli and visual stimuli are in different positions. Thus we cannot exclude the possibility that the difference between the auditory task and the visual task may not completely depend on the modality (i.e., visual Vs auditory) difference. We wanted to solve this problem by placing auditory and visual stimuli in the same position. We also use nose-poking to measure the reaction time in which a rat responds to stimuli. By using this behavioral system, we were able to confirm amodal recognition of space which means that a rat can respond to a different modality (visual or auditory) if the stimuli is in the same position and previously reported in other systems. We also confirm multisensory enhancement is indeed observed in rats. These resuluts suggests that this new modified AVD system can be used to explore the molecular and cellular mechanisms underlying multisensory processing in rats. The other task we developed is a wheel running system in which a water-deprived
mouse is asked to run to obtain water in front because the wheel with
the pegs is turning to the other direction (Kitsukawa et al., Society
for Neuroscience 32nd Annual Meeting, 2002). The pegs can be changed
with various patterns as desired. The task required for the mouse
thus can be regarded to represent a non-procedural learning. We examined
a various areas of brains following to the change of the peg pattern.
Among the areas examined, we found marked c-Fos expression in the
striatum. The striatum, which is composed of projection neurons and
several distinguished types of interneurons, is known to play an important
role in a reward-based learning. The characterization of these subtypes
of interneurons has been progressed. However, their roles in behavioral
tasks have been little known. We hope our system combined with c-Fos
mapping technique reveals it. We are currently doing collaborative
work with Professor Ann Graybiel (MIT) to record with tetrode from
the striatum under the task (supported by US-Japan exchange program).
We hope our wheel running system combined with c-Fos mapping technique
and electrophysiology reveals molecular and cellular mechanisms under
the Wheel Running task. CNTF, a member of the IL-6 family, attracts quite attentions of developmental neuroscientists because it shows various effects on neurons and glial cells. CNTF knockout mice, however, only indicate moderate motor neuron deficiency in the adult, but no apparent phenotype in the development. In order to explore the function of the IL-6 family, we extensively examined the expression of members of the family and their receptors and found the specific expression of CNTF in the embryonic pineal glands and eyes. This was to our knowledge for the first time to show a clear expression of CNTF in the development. Next question to be asked is the functional role of CNTF in the pineal development. Because of no reported phenotype of CNTF knockout mice in development, it has been difficult to know the developmental roles of CNTF if any. In fact, we also found there seemed no apparent difference in the pineal development of CNTF knockout mice compared to that in wild type mice. This is presumably because the CNTF gene knockout is largely compensated by other CNTF-like factors. We therefore studied cultured pineal organs in rodents to ask if there
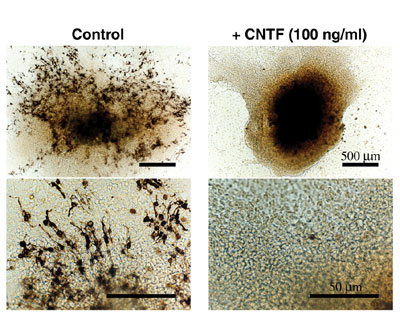
were any effects of CNTF on it following to a previous study that
shows that cultured neonatal pineal organs develop photoreceptor-like
cells (Araki, 1992). We found that CNTF inhibits photoreceptor-like
cells (Fig. 3, Hata et al., Dev Brain Res., 2003). Our observation
raises an interesting possibility that CNTF plays a critical role
in the pineal development to suppress a certain phenotype such as
photoreceptors. It also raises a question what roles CNTF play in
developing pineal glands if it play any roles in the development.
CNTF has been studied for nearly thirty years and still the role in
vivo remains to be studied. We hope that our study gives a clue to
solve this long standing question.
|
|||||||
Publication List: Tochitani, S., Hashikawa, T. and Yamamori, T. (2003) Expression of occ1 mRNA in the visual cortex during postnatal development in macaques. Neurosci. Lett. 337, 114-116 Suga, K., Yamamori, T. and Akagawa, K. (2003) Identification of Carboxyl-Terminal Membrane-Anchoring Region of HPC-1/Syntaxin 1A by Substituted-Cysteine-Accessibility Method and Monoclonal Antibody. J. Biolchem. 133, 325-334 Nakayana, T., Mikoshiba, K., Yamamori, T. and Akazawa, K. (2003) Expression of syntaxin 1C, an alternive splice variant of HPC-1/syntaxin 1A, is activaed by phorbol-ester stimulaton in astrglioma. FEBS Lett. 536, 209-214 Tochitani, S., Hashikawa, T. and Yamamori, T. (2003) Occ1 mRNA expression reveals a characteristic feature in the hippocampal CA2 field of adult mazaques. Neuroscience Lett. 346, 105-108 Hata, K., Araki, M., and Yamamori, T. (2003) Ciliary neurotrophic factor inhibits differentiation of photoreceptor-like cells in rat pineal glands in vitro. Dev. Brain Res. 143, 179-187 |